Synthesis and Characterization of Novel Potassium Ethane-1,2-diylbis((2-bromobenzyl) carbamodithioate) Complexes, Along with an Investigation of their Biological Activity.
1Department of Chemistry, College of Education for Women, University of Anbar, Iraq.
2Department of Chemistry, College of Science, University of Anbar, Iraq.
Corresponding Author E-mail:edw.aeedchemistry@uoanbar.edu.iq
DOI : http://dx.doi.org/10.13005/ojc/410516
ABSTRACT:In this study, the synthesized ligand and its complexes demonstrated notable antibacterial activity against several pathogenic bacteria. The Spectroscopic and analytical characterisation of a group of metallic complexes (II) belonging to the ligand dithiocarbamate were prepared, the ligand was prepared from three stepes. Follow: I. preparation of Schiff base by reaction Ethelinediamin with Bromobenzaldehyde. II. Reaction the product's Schiff base with Soduimborohydrate to yield Secondary Amin. III. Theproduct secondary amin reacted with carbondisulfide to producted ligand, which reacted with metal ion to yield dithiocarbamate complexes. The spatial shape around the ionic centers of the complexes was determined using a number of physic-chemical and spectroscopic techniques, including. UV-Vis, 1H-NMR, FTIR, mass spectrometers, melting points, and conductivity. This analytical and spectroscopic data showed that the MnII, FeII, CoII, NiII, CuII, and ZnII ions had a tetrahedral shape and square planar geometry with Ni(II) complexes. The antibacterial activity of ligands and complexes was evaluated with two types of bacteria: Staphylococcus aureus (gram-positive) and Escherichia coli (gram-negative). Zone of inhibition assays further supported these findings showing clear inhibition Zone of up to 15mm around the complexes. May be its disrupt bacterial cell membrane integrity and inhibit essential complexesas effective antibacterial agent.
KEYWORDS:Antibacterial; Complexes Characterization; Dithiocarbamate complexes; Macrocyclic Inorganic chemistry; Octahedral; Tetrahedral
Introduction
The journey for unused antibacterial specialists has ended up progressively basic in light of rising antimicrobial resistance.1 This ponder centers on the union and characterization of a novel ligand, potassium ethane-1,2-diylbis((2-bromobenzyl)carbamodithioate) complexes. These complexes are balanced to improve our understanding of metal-ligand intuitive whereas illustrating potential helpful applications.2 Characterization of the synthesized complexes will be conducted utilizing different methods, counting infrared spectroscopy, and atomic attractive reverberation. These strategies will offer assistance explain the auxiliary highlights and affirm the effective arrangement of the complexes.3,4 Ligands with interesting auxiliary properties can altogether impact the natural exercises of metal complexes. The joining of metal ion with dithiocarbamate ligand is especially vital; it is known for its differing pharmacological properties, counting antimicrobial impacts. By investigating the interaction between this ligand and metal ions, we point to create complexes that display upgraded steadiness and bioactivity.5,6 By giving a union, characterization, and organic assessment of these complexes, this inquire about points to contribute to the progressing explore for successful antimicrobial operators, advertising a establishment for future advancements in restorative chemistry.7,8 The knowledge into the natural assessment of these novel complexes, this research points to contribute to the progressing rummage around for compelling antimicrobial operators, advertising a establishment for future advancements in therapeutic chemistry.9,10
Material and Exprimental
Materials
2-Bromobenzaldehyde( BrC6H4CHO), Ethylenediamine(C2H8N2), potassium hydroxide (KOH), methanol (CH3OH), carbon disulphide (CS2), and dimethyl sulfoxide (DMSO) were bought from B.D.H, Merck, Fluke, and Sigma-Aldrich, and used without additional purification. Solvents were purified by distillation.
Synthesis of the Schiff base
Preparation of the Schiff-base (N1Z,N2E)-N1,N2-bis(2-bromobenzylidene)ethane-1,2-diamine was prepared by dissolving (3.07 g,16.6 mmol) of 2-Bromobenzaldehyde in (10 ml )of methanol and (3-4) drops of HBr, then (0.50 g, 8.31 mmol) of Ethylenediamine was added to the solution. Then the mixture was left for the reflux process for (3) hours at a temperature of (76-78 0C). After the reaction was completed, the mixture was left to cool and filtered, then the solvent was evaporated under vacuum pressure, giving a white precipitate with a weight of 2.94 g, (89.9%), m.p = 95-97 0C.
Preparation of secondary amine
The compound Schiff base was prepared dissolved (0.42 g, 1.06 mmol) from a Schiff base . In 40 mL (1:1) of the dichloromethane/methanol mixture, was added (0.24 g, 6.31 mmol) from NaBH4 to the solution, in an ice bath, leave the mixture to stir For a period of (12 hours), the organic layer was separated by adding (200 ml) water to the additive solution dichloromethane and the organic layer was washed with water four times (50 x 4 ml) and then dried With MgCO3, After the filtering and removing of the solvent by evaporation, the result is washed with diethyl ether. The product yellow solid.Yield: 0.30 g (71.4%). m.p = 87-89 ºC.
Synthesis of ligand [L]
The ligand Potassium ethan-1,2-diylbis((2-bromobenzyl)carbamodithioate was prepared by dissolving (0.50 g,1.25 mmol) of the aforementioned secondary amine in (20 ml) of methanol, and adding to it (0.14 g, 2.5 mmol) dissolved in (5 ml) of methanol, with continuous stirring for 15 minutes after which the solution was cooled in an ice bath to add (0.18 g, 2.48 mmol) of CS2 was left to stir continuously for( 2-3) hours to give the precipitate dithiocarbamate salt formed after filtering and washing with methanol and ether and then drying it under vacuum pressure to give a white precipitate by weight 0.63 g (90%), m.p = 100-102 ºC .
General synthesis of macrocyclic-based complexes
The complexes were prepared from the reaction of one mole of the potassium dithiocarbamate (K-DTC) salt against one mole of the mineral salt as a central ion to have the ratio (1: 1), as the metals used to prepare the cyclic complexes are Mn(II), Fe(II), Co(II), Cu(II) , Ni(II), and Zn(II). The mixture containing the ligand salt and the metal and their solvents are stirred for 24 hours, and distilled water is added if necessary to complete the sedimentation. We obtain the precipitate by filtering, washing it with solvent, and drying it to give complexes of different colors according to the type of metal to give the ring complex.
Determination of bacteriological activity
The biological activity of the ligand and selected complexes, [Mn(L)]2, [Fe(L)]2, [Co(L)]2, [Ni(L)]2, [Cu(L)]2, and [Zn (L)]2, was studied on two different types of bacteria.
Some chemicals were tested and a solvent (DMSO) was employed. 50, 100, and 150 mg/ml The 50 and 100 mg/ml concentrations were chosen for the biological action. The bioactivities were examined by agar well diffusion. The wells were dug in the media with the help of a sterile metallic borer with centers of at least 24 mm. The concentration of (50 and 100) mg/mL in DMSO was applied to each well. The plates were incubated at 37°C for 24 hours. Zones indicating complete inhibition were used to determine the activity (mm). Increasing the width of the inhibition signifies an increase in the biological efficacy of the produced compounds compared to antibiotics’ (Ghoto, Khuhawar and Jahangir, 2019).
Phyasical Mesuerment
An Electro-Thermal Stuart melting point apparatus was used to determine the melting points of compounds. On a Shimadzu and Bruker (FTIR)–8400S spectrophotometer, Fourier transform infrared spectra (FTIR) were recorded in the range (4000–400 cm–1). As KBr disks, spectra were obtained. The electronic spectra of ligands and their complexes were calculated in DMSO solutions with a concentration of (10-3-10-5 M) using a JASCO -V- 650 UV-Visible Spectrophotometer. On a Bruker 500MHz SWISS TS AG, 1H NMR spectra were recorded. an NOV AA (F.A.A) 350G atomic absorption spectrophotometer was used to determine the metal content of complexes. Jenway conductivity meter model 4070 was used to record conductivity measurements of the complexes at (25 oC) for (10-3 mole L-1) solution of the samples in DMSO.
Resulte and Disscution
In this study, the chemical compounds (Schiff’s base), (secondary amine) and (ligand salt) were prepared, as it included the preparation of a Schiff base from the reaction of the primary amine with 2-bromobenzaldhyde with the presence of methanol solvent, after the formation of a Schiff base that was dissolved in The solvents of ethanol and dichloromethane were then added NaBH4 to give the product, which is the secondary amine, where the amine was dissolved, then carbon disulfide (CS2) and the base (KOH) were added, which gave the ligand salt in a good proportion. The metal ratio is 1:1. In this reaction, the metal ion plays a large and prominent role in the self-preparation of metal complexes. The product gave two general formulas for the complexes, which are the tetrahedral shape and the octahedral shape as shown in (Scheme 1). The prepared organic compounds were diagnosed by mass spectrometry, spectrometry. Infrared, melting point, molar conductivity, nuclear magnetic resonance spectroscopy, and UV-visible.
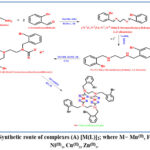 |
Scheme 1: Synthetic route of complexes (A) [M(L)]2; where M= Mn(II), Fe(II), Co(II),, Ni(II),, Cu(II),, Zn(II),. |
Table 1: Colour, yield, melting point, and molar conductivity of Complexes.
| Complexes | Weight of metal salt (g) | Weight of complex(g) | M.Wt | Yield (%) | Colour | m.p. ºC | ƹM(Ω cm-1 2 mol) in DMSO |
| L | 0.5 | 0.63 | 626.58 | 90 | Off-white | 100* | 0.52 |
| [Mn(L)]2 | 0.4 | 0.99 | 1302.76 | 82 | Dark brown | 111* | 0.39 |
| [Fe(L)]2 | 0.42 | 0.89 | 1304.58 | 74 | Dark red | 140* | 0.98 |
| [Co(L)]2 | 0.23 | 0.36 | 1310.75 | 50 | green | 95* | 0.3 |
| [Ni(L)]2 | 0.24 | 0.46 | 1382.34 | 65 | green | 255* | 0.37 |
| [Cu(L)]2 | 0.11 | 0.19 | 1392.04 | 55 | Dark green | 103* | 2.03 |
| [Zn(L)]2 | 0.11 | 0.29 | 1323.65 | 85 | off-white | 180* | 0.56 |
FTIR spectra
Herein are presented the FTIR spectra of L (Figure.1) and its complexes dipected in table (2). Two bands at 1632-1585 cm-1 in the L FTIR spectrum referred to v(C=N) and ν(C=C) of the pyridyl ring. The strong band in the range 1460-1500 cm-1 is explained by the ν(N-CS2) stretching vibration. Compared to the unbound ligand at 1463 cm-1, this band demonstrates an increase in the carbon-nitrogen double bond character. the bands at 1023 and 944 cm-1 in the ligand Anisobidentate chelation mechanism of the ligand was shown by these bands.11,12 Also, the complexes of the first ligand appeared new absorption bundles linked to the ν (M-S) bond, namely ν (Mn-S) (439) cm-1, ν (Fe-S) (439) cm-1, As in ν (Co-S), (424)cm-1, ν (Cu-S) (432) cm-1, ν (Zn-S) (408) cm-1,13 Table (2).
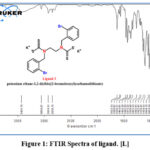 |
Figure 1: FTIR Spectra of ligand. [L]. |
Table 2: FTIR spectral data (wave number) cm-1 of L3 and its complexes.
| Comp. | νar(C=C) | ν(N-CS2) | ν(C-N) | νas,s(CS2) | (M-S) ν |
| L | 1566 | 1463 | 1265 | 1023, 944 | _ |
| [Mn(L) 2 ] | 1512 | 1423 | 1342 | 1002,860 | 439 |
| [Fe(L)]2 | 1566 | 1435 | 1346 | 1022,983 | 439 |
| [Co(L)]2 | 1562 | 1438 | 1323 | 1002,891 | 424 |
| [Ni(L)]2 | 1566 | 1485 | 1346 | 1026,948 | 416 |
| [Cu(L)]2 | 1516 | 1423 | 1369 | 1045,960 | 432 |
| [Zn(L)]2 | 1516 | 1419 | 1319 | 1026 | 408 |
Electronic_ spectra and magnetic moments
The UV-Vis spectra of the Mn+2 complex showed an absorption peak at (518) nm, (19305) cm-1, which goes back to 6A1g → 4T1g (G), in which this transition is attributed to the tetrahedral.14,15 As for the Fe+2 complex, an absorption peak of (554) nm, (18050) cm-1 appeared, with an electronic transition of the d-d type, which belongs to the transition 5E2→ 5T2, which is attributed to this transition to the tetrahedral shape.16,17 For the Co+2 complex, a peak appeared at (641) nm, (15600) cm-1, which is due to the transition 4T1g(F) → 4T2g(P), which is attributed to this transition to the tetrahedral shape.12 For the Ni+2 complex, the spectrum shows two main peaks at(640) nm, (15625) cm-1 which refers to the transition 1A1g → 1A2g, which is attributed to this transition to the square planar shape.15 For the Cu+2 complex, a peak at (658) nm, (975) cm-1 is due to the transition 2B1g→2A1g, which is attributed to this transition to the tetrahedral shape.15 For the Zn+2 complex, a peak appeared at (425) nm, (23529) cm-1 which is C.T. The transition is attributed to the tetrahedral shape18,19see Table (3).
Table 3: UV-Vis Spectral data of L in DMSO solutions
| Comp. | Concen.
mol/L |
Band
Positio n λnm |
Wave
number (cm-1) |
Assignment | Suggested
geometry |
| [Mn(L)]2 | 1×10-5
|
262
306 518 |
38167
32679 19305 |
Intra-ligand π → π*
n → π* 6A1g(F) → 4T1g(G) |
Tetrahedral |
| Fe(L)]2 | 1×10-5
1×10-3 |
275
315 554 |
17391
31746 18050 |
Intra-ligand π → π*
n → π* 5E2D) →5T2 (D) |
Tetrahedral |
| [Co(L)]2 | 1×10-5
1×10-3 |
269
325 423 641 |
37174
30769 23640 15600 |
Intra-ligand π → π*
n → π* C.T 4T1g(F) → 4T2g(P) |
Tetrahedral |
| [Ni(L)]2 | 1×10-5
1×10-3 |
259
394 640 |
38610
25380 15625 |
Intra-ligand π → π*
n → π* 1A1g → 1A2g |
square planar |
| [Cu(L)]2 | 1×10-5
1×10-3 |
260
401 658 |
38461
24937 15197 |
Intra-ligand π → π*
n → π* 2B1g→2A1g |
Tetrahedral |
| [Zn(L)]2 | 1×10-5 | 262
425 |
38167
23529 |
Intra-ligand π → π*
C.T |
Tetrahedral |
1H NMR SPECTRUM
1H NMR spectrum for ligand (L)
The structural formula of the prepared compound was confirmed using 1H-NMR proton nuclear magnetic resonance spectroscopy and using the solvent [DMSO-d6], which gave further evidence for the correctness of the product formation through the disappearance of the single signal at the region [δ = 4.6ppm] of a proton. The secondary amine group and the single sign appearing at the range [δ = 3.84ppm, (2H, s), CH2 alip.], is due to (C6,6`-H), Belonging to the protons of the two symmetric aliphatic groups in the compound, and a single signal appeared at the range [δ = 4.24ppm, (2H, s) CH2 benzyl], is due to (C5,5`-H), belonging to the two proton groups associated with the homologous aromatic ring in the compound, and the multiple signals of the aromatic ring protons at the range [δ = 7.89-8.21 ppm, (4H, m), Ar-H], is due to (C1,1`,2, 2`, 3, 3`, 4, 4`-H), which show a clear correspondence with the values in The scientific literature.20,21
Table (4) shows the chemical displacement of the prepared compound diagnosed by the NMR spectrum measured in ppm. Shown in Figure (2).
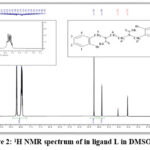 |
Figure 2: 1H NMR spectrum of in ligand L in DMSO-d6 |
1H NMR spectrum of [Zn(L)]2
The structural formula of the prepared compound was confirmed using 1H-NMR proton nuclear magnetic resonance spectroscopy and using the solvent [DMS0-d6], which gave further evidence for the correctness of the product formation through the observed change in the locations of the chemical displacements of the protons and the increase in the number of protons it showed. The spectrum, showing a single signal at the range (δ = 3.76ppm, (4H, s), CH2 alip.), is due to (C6,6`-H) Belonging to the protons of the symmetric aliphatic groups in the compound, and a single signal at the range [δ = 4.68ppm, (4H, s) , CH2 benzyl], is due to (C5,5`-H), Belonging to the protons of the two groups associated with the symmetric aromatic ring in the compound, and the multiple signals of the aromatic ring protons at the range [δ = 7.81 – 8.42 ppm, (8H, m), Ar-H], is due to (C1,1`,2, 2`, 3, 3`, 4, 4`-H), which show a clear correspondence with the values in the literature Scientific.22,23 Table (4) shows the chemical displacement of the prepared compound diagnosed by the NMR spectrum measured in ppm. shown in Figure (3).
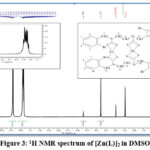 |
Figure 3: 1H NMR spectrum of [Zn(L)]2 in DMSO-d6 |
Table 4: 1H-NMR data for ligand and Complex in DMSO-d6
| Compound | Functional group | δ =(ppm) |
|
Free ligand L |
(C6,6`-H) | 3.84 (2H, s), |
| (C5, 5` -H) | 4.24 (2H, s) | |
| (C1,1`,2, 2`, 3, 3`, 4, 4`-H) aromatic ring | 7.89-8.21 (4H, m) | |
| DMSO | 2.51 | |
| HDO | 3.33 | |
| [Zn(L)] 2 | (C6,6`-H) | 3.76 (4H, s) |
| (C5, 5` -H) | 4.68ppm, (4H, s) | |
| (C1,1`,2, 2`, 3, 3`, 4, 4`-H) aromatic ring | 7.81 – 8.42 (8H, m) | |
| DMSO | 2.51 | |
| HDO | 3.31 |
Mass Spectrum
Mass spectrum of Schiff base
The electrospray (+) mass spectrum of Schiff base is presented in Figure (4). A series of fragments related to ligand structure can be seen in the spectrum. the parent ion peak for the ligand at m/z = 394.95, which corresponds to (M)+ ( percent ) for C16H14Br2N2; requires =394.16, as well as other fragments, relative abundance, and fragmentation pattern shown in Table (5).
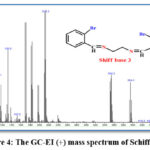 |
Figure 4: The GC-EI (+) mass spectrum of Schiff base. |
Table 5: GC-Mass data of the Schiff base
| Fragment | Mass/charge(m/z) |
| [M]= [C16H18Br2N2+H] | 395.2 |
| [C9H11BrN] + | 213.1 |
| [[C7H6Br] + | 169.1 |
| [C4H4Br]+ | 132.2 |
| [C4H11N2]. | 88.1 |
Mass spectrum of secondary amine
The electrospray (+) mass spectrum of secondary amine. is presented in Figure (5). A series of fragments related to ligand structure can be seen in the spectrum. The ligand’s parent ion peak is found at m/z = 395.2, which corresponds to (M)+ ( percent ) for C16H18Br2N2; requires = 398.14, as well as other fragments, their relative abundance, and fragmentation pattern are shown in Table (6).
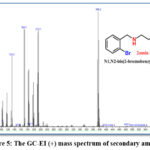 |
Figure 5: The GC-EI (+) mass spectrum of secondary amine. |
Table 6: GC-Mass data of the secondary amine
| Fragment | Mass/charge(m/z) |
| [M]= [C16H18Br2N2+H] | 395.2 |
| [C9H11BrN] + | 213.1 |
| [[C7H6Br] + | 169.1 |
| [C4H4Br]+ | 132.2 |
| [C4H11N2]. | 88.1 |
Mass spectrum of free ligand [L]
The electrospray (+) mass spectrum of KL is presented in Figure (6). A series of fragments related to ligand structure can be seen in the spectrum. The ligand parent ion peak is found at m/z =625.4, which corresponds to (M)+ ( percent ) for C18H16Br2K2N2S4; requires =626.58, as well as other fragments and their relative abundance and fragmentation pattern are shown in Table (7).
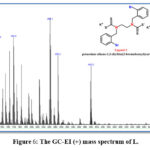 |
Figure 6: The GC-EI (+) mass spectrum of L. |
Table 7: GC-Mass data of the ligand KL
| Fragment | Mass/charge (m/z) |
| [M]= [C18H16Br2K2N2S4 +H] | 625.4 |
| [C12H13 BrKN2S3]+ | 400.2 |
| [C11H14 BrN2S2] + | 288.2 |
| [C15H21N2S] + | 262.1 |
| [C3H7 Br N] + | 138.2 |
| [C4H11N2]• | 89.2 |
| [CH3S] + | 50.1 |
Antibacterial Activity
The results studied at concentrations of 50 mg/ml and one 100 mg/ml and compared with anti-inflammatory such as Ciprofloxacin and Levofloxacin standards showed: that ligand shows antimicrobial activity against Escherichia coli (E. coli) and Staphylococcus aureus. However, the free ligand showed higher antimicrobial activity against Escherichia coli (E. coli), of the metal complexes compared with some of the measured antibiotics, see Table (8, 9). Furthermore, the [Cu(L)]2 complexes showed higher activity against Staphylococcus aureus compared to the free ligand at concentration (100,50 ) indicated that significant inhibitory activity was observed at concentration (100 mg/ml) for ([Mn(L)]2, [Fe(L)]2, [Co(L)2], [Ni(L)]2, [Zn(L)]2 compounds, while, [Mn(L)]2 , [Fe(L)]2, [Ni(L)]2,[ and [Zn(L)]2 at its concentration (50 mg/ml) showed moderate inhibiting activity among most bacteria.24,25
Table 8: Results of an antibacterial study of complexes and prepared ligand
| No. | Complexes | Conc.mg/ml | staphylococcus
aureus |
Escherichia coli |
| 1 | KL | 50
100 |
20
25 |
27
32 |
| 2 | [Mn(L)]2 | 50
100 |
12
15 |
11
13 |
| 3 | [Fe(L)]2 | 50
100 |
12
15 |
10
11 |
| 4 | [Co(L)]2 | 50
100 |
22
23 |
16
18 |
| 5 | [Ni(L)]2 | 50
100 |
16
22 |
12
17 |
| 6 | [Cu(L)]2 | 50
100 |
32
32 |
24
25 |
| 7 | [Zn(L)]2 | 50
100 |
17
25 |
14
16 |
Table 9: The inhibitory activity of a number of control factors (antibiotics) on the growth of a number of positive and negative bacteria.
| No. | Complexes | Conc.mg/ml | staphylococcus
aureus |
Escherichia coli |
| 1 | Ciprofloxacin | 50
100 |
—
— |
25
26 |
| 2 | Levofloxacin | 50
100 |
—
— |
25
26 |
Conclusion
In this research, the free ligand and its interaction with ions of metallic elements were prepared and diagnosed and the complexes were given the stereoscopic shape through spectroscopic and analytical measurements that proved the formation of two types of shapes, the tetrahedral shape and the square planar around the center of the metal. The complexes’ biological activity was tested on a variety of bacterial strains. Pathogens such as Escherichia coli (E. coli) and Staphylococcus aureus were particularly vulnerable to the complexes. After examining the in vitro effect of metal complexes on experimental animals and clinical trials, the current study suggested that complexes will be employed as promising candidates for further biological evaluation.
Acknowledgement
We want to express my sincere gratitude to the Departments of Chemistry and Biology at University of Anbar for its invaluable support and resources throughout our research.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
References
- Yousif, E.I.; Ahmed, R.M.; Hasan, H.A.; Al-Fahdawi, A.S.; Al-Jeboori, M.J. Iranian Journal of Science and Technology, Transactions A: Science, 2017, 41, 103-109.
CrossRef - Breijyeh, Z.; Karaman, R.; Antibiotics (Basel, Switzerland), 2023,12 (3), 628.
CrossRef - Bakheit, A.H.; Al-Salahi, R,; Ghabbour, H. A.; Ali, E.A.; AlRuqi, O. S.; Mostafa, G. A .E.; Crystals; 2023, 13(7),1088.
CrossRef - Alsalihi, I.I.; Al-khafaji, Y.F,; Al-fahdawi, A.S.; Atiyah E M.; Bulletin of the Chemical Society of Ethiopia; 2022, 36(4), 791-799.
CrossRef - Liu, R.; Cui, J.; Ding, T.; Liu, Y.; Liang, H.; Molecules (Basel, Switzerland), 2022, 27(23), 8393.
CrossRef - Juliet, A.A.; Hameed, A.T.; Ali, F.F.; HIV Nursing. 2022. , 22(2):1631-4.
- Abdulqader, I. A.; Al-Fahdawi , A.S.; fadhel Ali, F.; Journal of Survey in Fisheries Sciences. 2023, 10(3S):2890-905.
- Saleh, S.; Al-Timari, U.; Al-Fahdawi , A.; El-Khatatneh, N.A.; Mahendra, M.; Al-Ghorbani, M.; Oriental J. Chem. 2017, 33(6):2713-9.
CrossRef - Yan, Y.; Li, X.; Zhang, C.; Lv, L.; Gao, B.; Li, M.; Antibiotics (Basel, Switzerland), 2021, 10(3), 318.
CrossRef - Muteeb, G.; Rehman, M.T.; Shahwan, M.; Aatif, M.; Pharmaceuticals. 2023, 16(11):1615.
CrossRef - Ghoto, S.A.; Khuhawar, M.Y.; Jahangir, T.M.; Analytical sciences. 2019, Jun;35(6):631-7.
CrossRef - Al-Obeidi, F.Z.; Al-Fahdawi , A.S.; Al-Jeboori, M.J.; J. Global Pharma Technol. 2018;10(03):699-710
- Abdullah, S.A.; Othman, E.A.; Al-Fahdawi, A.S.; Samarra Journal of Pure and Applied Science. 2020, 2(3).
- Al-Mohammadi, N.A.; Al-Fahdawi , A.S.; Al-Janabi, S.S.; Iraqi J.of Science. 2021, Jan 30:1-5.
CrossRef - Al-fahdawi, A.S.; Al-Sorchee, S.M.; Saleh, S.E.; Saleh, M.M; Egyptian Journal of Chemistry, 2021, 64(6), pp. 2879–2888
- Salih, B.D.; Dalaf, A.H.; Alheety, M.A.; Rashed, W.M.; Abdullah, I.Q.; Materials Today: Proceedings. 2021. Jan 1;43:869-74.
CrossRef - Al-Fahdawi, A. S.; Herman, P.; Al- Jeboori, M. J.; Asian Journal of Green Chemistry ; 2025 ,9 ,1 , 32-42
- Andrew, F.P.; Ajibade, P.A.; International Journal of Electrochemical Science. 2019, Aug 1;14(8):7062-75.
CrossRef - Al-fahdawi, A.S.; Mohamed, S.K.; Potgeiter, H; Alfutimie, A. ; Bulletin of the Chemical Society of Ethiopia; 2024, 38(6):1583-93
CrossRef - Yousif, E.I.; Hasan, H.A.; Ibn AL-Haitham Journal For Pure and Applied Science. 2017 Jun 11;30(1):73-87.
CrossRef - Sultan, J.S.; Alsalihi, E.I.; Al-Fahdawi, A.S.; Jabar, A.; Egyptian Journal of Chemistry, 2020, 63(1), pp. 325–335
CrossRef - Hussain, S.; Zahid, Z.; Shahid, M,; Abid M.A.; Arabian Journal for Science and Engineering. 2019. Jul 1;44:6423-34.
CrossRef - Al-Sorchee, S.M.; Ismail, T.F.; Mohammed, I.M.; Al-fahdawi, A.S.; Biochemical and Cellular Archives, 2020, 20(1), pp. 373–382
- Al-Obaidy, G.S.; Al-Dulymi, Hk.Hk.; Abed, M.A.; Sciences; 2024, 22(2):1082-93.
CrossRef - Singh, R.V.; Dwivedi, R.; Joshi, S.C.; Transition Metal Chemistry. 2004 Feb;29:70-4.
CrossRef
Accepted on: 16 Oct 205
Second Review by: Dr. K. Koteswara Rao
Final Approval by: Dr. Ioana Stanciu

















