Biogenic Synthesis, Characterization and Antibacterial Activity of Silver Nanoparticles Mediated by Ziziphus Spina Christi Leaf Extract
Department of Botany and Microbiology, College of Science, King Saud University, Riyadh, Saudi Arabia
Corresponding Author E-mail:ssalmen@ksu.eda.sa
DOI : http://dx.doi.org/10.13005/ojc/410531
ABSTRACT:The study of biogenic nanoparticles (NP) as antibacterial agents is a novel and rapidly developing area of medical biotechnology. The current work used Zizyphus spina-christi aqueous leaf extract to reduce the silver ions in the silver nitrate, thereby resulting in the production of nanoparticles of silver through biosynthesis. The fabrication and characterization of nanoparticles biosynthesized using leaf extracts of Zizyphus spina-christi were observed using a variety of techniques including, UV visible spectroscopy, EDS, SEM, TEM, and FTIR. AgNPs in diameters ranging from 5 to 30 nm were produced. The FTIR spectrum identified certain functional compounds responsible for the biosynthesis of these AgNPs. The agar disc diffusion technique was used to assess the antibacterial effects of AgNPs biogenically synthesized using leaf extract of Zizyphus spina christi confirmed antibacterial effect against bacteria that belong to both Gram- negative and Gram-positive (but were inactive against Salmonella typhimurium.) AgNPs fabricated using this plant extract can here are potentially of use in medicine as antibacterial agents.
KEYWORDS:Antibacterial activity; Bacteria; Biogenic synthesis; Nanoparticles; Zizyphus spina-christi
Introduction
Nanotechnology has rapidly expanded during the past ten years worldwide at the level of research, industrial activity, and product development in order to create environmentally friendly nanomaterials for use, in for example, water treatment, medicine, and solar energy conversion.1-8 Metal particles varying between 1-100 nm in diameter are now the object of significant research because of their unique chemical, optical, electrical, adjustable hydrophilic–hydrophobic balance and catalytic features, as well as their significant ratio of surface area to volume and strong antibacterial effects1. Nanomaterials have been fabricated from various metals, including silver, gold, magnesium, alginate, titanium, zinc and copper.9-11 Among the most widely used nanoparticles, sliver nanoparticles have been reported to have significant antibacterial activity12. Numerous techniques can be employed to synthesize (NPs), such as chemical reduction, sono-electrochemical techniques, microwave irradiation and ultraviolet. Nowadays, biogenic fabrication, also known as green synthesis, is advantageous over conventional methods in being safer, cheaper, and more environmentally friendly.5,6,13-15 Silver nanoparticles synthesized using biological methods have antibacterial effects,16,17 antioxidant, and anti-platelet activities18; anti-angiogenesis19; anti-viral activities20 and anti-inflammatory effect.21 Several plants extract such as A. indica (Neeem), M. oleifera (Horseradish), C. colocynthis (Bitter Apple) and T. foenum-graecum (Fenugreek) etc. have been positively used for the biogenic synthesis of metallic nanoparticles.22, 23
Ziziphus spina-christi or Sidr tree, is an evergreen plant which is indigenous to the majority of nations in southern western Asia and Africa countries.24 Along with several recently imported noxious species of weeds, It is believed that Z. spina-christi is one of the few Saudi Arabian native tree species still in existence.25,26 Sidr leaves are used in traditional treatments as a demulcent, mouth wash, anodyne, depurative, emollient, astringent, for toothaches, and stomachic Zizyphus spina-christi leaf extract has been shown to contain flavonoids, tannins, phenolic, saponins and alkaloids, all of which are powerful agents of reduction because of their many OH-groups which are responsible for their antioxidant and antimicrobial activity.27,28 In this study, an aqueous leaf extract of Zizyphus spina christi L was applied to biosynthesize nanoparticles of silver by reducing the silver ions existing in the silver nitrate solution. After characterization, antibacterial effects of the biosynthesized AgNPs were investigated and assessed against a number of important pathogenic bacteria.
Materials and Methods
Materials
Zizyphus spina christi (Sidr) leaves were obtained from the Riyadh region, Kingdom of Saudi Arabia. Nutrient agar and both Mueller Hinton Broth and Mueller Hinton Agar (Sigma–Aldrich) were used as growth media. AgNO3 (1mM, Sigma–Aldrich) was prepared using double distilled water and stored in refrigerator at 4°C.
Preparation of Zizyphus spina christi leaf extract
The obtained leaves were cleaned several times with tap water and dried at 25°C in the shade, before being powdered in a grinder. In order to prepare the extract, 10 grams of finely ground Zizyphus spina christi L leaf was added to 100 milliliters of deionized water, which was then heated for 20 minutes, filter passed through filter paper (Whatman No. 1) and stored at 4°C.
Biogenic synthesis of AgNPs
AgNPs were produced by the biosynthesis approach. In order to reduce the Ag+ ions, 1 mL from leaf extract of Zizyphus spina Christi was added to 100 mL of an aqueous solution of AgNO3 (1 mM) and stirred at 60°C for 1 h. To confirm the formation of AgNPs, a dark brown color alteration was detected, indicating the synthesis of AgNPs. The AgNPs were then separated from the mixture using centrifugation for 20 min at 15,000 rpm, followed by drying the nanoparticles at 40 °C.
Characterization of AgNPs
UV visible spectroscopy was carried out between 200 and 800 nm in wavelength by a UV/visible Ultrospec 2100 Pro spectrophotometer (Biochrom, UK). EDS analysis was achieved using an Altima IV device (Regaku, Japan) as previously described29. Scanning electron microscopy (SEM) analysis of AgNPs was conducted using a JEOL microscope (JSM-6380 LA), while FT-IR Spectra was carried out using (Thermo Scientific Smart iTR™) for both AgNPs and Zizyphus spina christi leaf extract in order to monitor the existence of bioactive compounds responsible for NPs formation. The size of dimeters and shape of the AgNPs were observed via using a JEOL microscope (JEM-1011).
Antibacterial properties of AgNPs
Bacterial cultures were diluted with phosphate buffered saline to obtain 0.5 McFarland standards and inoculated on Petri dishes contained Mueller Hinton Agar. Sterile disks of paper were then saturated with AgNPs, put on the swabbed plate surface, and then cultured at 37°C for 18 to 24 hours. Areas of inhibitory around the paper disks were then measured in millimeters in order to assess antibacterial activity.
Statistical analysis
Statistical analysis was implemented using ANOVA, and results presented as average ± SD. P-value of less than 0.05 was considered statistical significance.
Results and Discussion
Ultraviolet‑visible spectroscopy
The first characterization of AgNPs synthesized using the aqueous extract of Z. spina-Christi L was carried out by UV-vis spectroscopy. AgNPs were prepared using Z. spina-Christi leaf extract with solution of AgNO3 (1 mM) at 25°C via Ag+ reduction to Ag0. One hour after the reaction was carried out, the colorless mixture changed to a dark brown, suggesting the formation of AgNPs. The change in colour of the sidr leaf extract shows the reducing capability of the plant extract associated with the fabrication of AgNPs30. The reaction kinetics were monitored in the range of 200–800 nm. Fig. 1 shows the spectrum of biosynthesized AgNPs by Sidr leaf extract. The extreme absorption wave of AgNPs at 400 nm illustrates the formation of AgNPs. The surface plasmon resonance is considered responsible for the extreme absorption peak of AgNPs at 400 nm and is attributed to the excitation of free electrons31. The observed colour change and maximum absorption wave at 400 nm are in agreement with a number of previous studies on extracts of plants such as A. vera, P. oleracea and C. dactylon32, I. Oblongifolia33 and A. fleurentiniorum.34
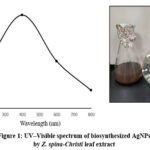 |
Figure 1: UV–Visible spectrum of biosynthesized AgNPs by Z. spina-Christi leaf extractClick here to View Figure |
FTIR analysis
The potential bioactive molecules and functional compounds in the plant filtrate responsible for the biosynthesis of AgNPs were determined recognized using FTIR. Fig. 2 (a, b) and table 1 show the FTIR spectra of Sidr leaf extract and the biogenically synthesized AgNPs. The extreme broadband at 3358 cm−1 results from the stretching of O–H groups35,36, while the strong second band in the FTIR profile, at 2974 cm−1 and 2880 cm−1 is related to the alkyl –C–H group37. The stretching vibration of (NH) C=O group is shown by a band at 1654 cm−1 current in carboxylic acids, ketones, quinones, and esters38,39. Other strong absorption wave at 1049 cm−1 may result from –C–O stretching vibrations of alcoholic compounds40. The FTIR spectrum of the AgNPs also revealed a similar FTIR spectral data derive from the plant filtrate. Finally, the results of FTIR confirmed the presence of O–H, –C–H –NH, (NH) C=O and –C–O group.
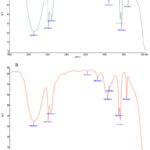 |
Figure 2: FTIR spectra of Z. spina Christi extract (A) and AgNPs fabricated using the Z. spina-Christi leaf extract (B).Click here to View Figure |
Table 1: Functional groups of AgNPs and Z. spina Christi extract fabricated by FTIR analysis.
| Treatment | Spectra | Functional group | Compound class |
| AgNPs | 3358 | O–H stretching | Alcohol |
| 2974 and 2880 | –C–H bending | alkyl | |
| 1654 | C=O bending | carboxylic acids, ketones quinones, and esters | |
| 1049 | –C–O stretching | Alcoholic compounds | |
| Z. spina Christi extract | 3358 | O–H stretching | Alcohol |
| 2974 and 2880 | –C–H bending | alkyl | |
| 1654 | C=O bending | carboxylic acids, ketones quinones, and esters | |
| 1049 | –C–O stretching | Alcoholic compounds |
EDS and SEM analysis
Fig.3 shows the EDS spectra of AgNPs, which showed significant indicators in the silver area at 3 keV, supporting the occurrence of elemental silver atoms; the EDS profile displayed the presence of silver 38.75%.in the nanoparticles. The appearance of signals for other elements such as carbon, oxygen and chlorine in the fig.3 may result from X-ray emissions of the peptides that are present in Z. spina-Christi leaf extract33. There are other explanations for the existence of identical signals, such as the potential attachment of molecules to AgNPs surfaces inducing the reduction of silver ions to elemental silver36. The above findings agree with the results of previous studies on a variety of plant extracts, including Indigofera oblongifolia33, Phoenix dactylifera seeds41, extract of Artemisia sieberi and Calotropis procera42. The SEM image of AgNPs in Fig.3 shows variously sized, irregularly formed AgNPs. AgNPs clusters, which can be linked with the accumulation of NPs during the sample mixture.
 |
Figure 3: SEM photograph and EDS of AgNPs produced using leaf extract of Sidr (Z. spina-Christi).Click here to View Figure |
Morphology and size of AgNPs
The shape and sizes of silver nanoparticles were investigated using TEM technique in this study. Fig. 4 shows a TEM photograph of nanoparticles produced using Sidr leaf extract as well as the size-range of the resultant nanoparticles; the AgNPs were circular in shape with dimensions among from 5 to 30 nm. Particles were well distributed and crystalline, with a pale organic cover. Small particles were also found to form minor, a finding which may result due to accumulation or improper capping. These results are consistent with previous findings, which generally reported a particle size of <50nm36,43, although. differences in the size of dimeters and forms of the nanoparticles fabricated have also been reported following the use of organic synthesis techniques.44,45
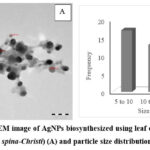 |
Figure 4: TEM image of AgNPs biosynthesized using leaf extract of Sidr (Z. spina-Christi) (A) and particle size distribution (B).Click here to View Figure |
Antibacterial effect of AgNPs
The antibacterial effect of AgNPs biogenically fabricated using leaf extract of Sidr plant was assessed against pathogenic microorganisms like the Gram-negative bacteria, E. coli and S. typhimurium and the Gram-positive bacteria, B. subtilis and S. aureus using the agar disc diffusion assay. Inhibition zones (mm) around each disc containing AgNPs are shown in Table 2. With the exception of S. typhimurium, all of the bacteria used showed significant susceptibility to AgNPs synthesized using Z. spina-Christi leaf extract. AgNPs’ antibacterial activity results from the disruption of the respiratory chain and thereby cell division, which leads to cell death. Additionally, it has been shown that the silver nanoparticles present inside bacterial cells release silver ions, which increases their bactericidal action.46,47
Table 2: Antibacterial activity of AgNPs on some pathogenic bacteria
| Name of bacteria | Inhibition zone (mm) | |
| AgNPs | Control | |
| S. aureus | 8± 0 | 0 |
| B. subtilis | 9±1.1 | 0 |
| S. typhimurium | 0 | 0 |
| E. coli | 10± 0.57 | 0 |
Conclusion
An easy, low-cost and an eco-friendly technique for the fabrication of AgNPs using the aqueous extract of Sidr (Z. spina-Christi) was developed in the present study. Characterization of AgNPs demonstrated that the particles of silver were circular with sizes among from 5 to 30 nm. AgNPs were shown to exhibit antimicrobial effects against some pathogenic microbes such as both Gram-positive species and Gram-negative species (with the exception of Salmonella typhimurium). AgNPs fabricated using the extract of Sidr plant potentially could be used as antibacterial agents in medicine.
Acknowledgement
This project was supported by Ongoing Research Funding program (ORF-2025-385), King Saud University, Riyadh, Saudi Arabia.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
References
- Whitesides, G.M. Small. 2005,1,172-179.
CrossRef - Filipponi, L.; Sutherland, D. Interdisciplinary NanoscienceCenter (iNANO). Denmark: Aarhus University; (2010).
- Asif, M.; Yasmin, R.; Asif, R.; Ambreen, A.; Madiha, M.; Umbreen, S. Dose-Response: An International Journal. 2022,2,1–11.
- Seku, K.; Gangapuram, B.R.; Pejjai, B.; Kadimpati, K.; Golla, N. J Nanostruct Chem. 2018,8,179–88.
CrossRef - Rai, M.; Yadav, A.; Gade, A. Biotechnol Adv. 2009,27,76-83.
CrossRef - Torabfam, M.; Yüce, M. Green Process Synth. 2020,9,283–293.
CrossRef - Shanmuganathan, R.; MubarakAli, D.; Prabakar, D.; Muthukumar, H.; Thajuddin, N.; Smita Kumar, S. Pugazhendhi, A. Env Sci Pollut Int. 2018, 25,10362–70.
CrossRef - Abel, S.; Tesfaye, J.L.; Shanmugam, R.; Dwarampudi, L.P.; Lamessa, G.; Nagaprasad, N.; Benti, M.; Krishnaraj, R. J Nanomater. 2021,6, 2021.
CrossRef - Ahmad, Z.; Pandey, R.; Sharma, S.; Khuller, G.K. Indian J Chest Dis Allied Sci. 2006,48,171-176.
- Gu, H.; Ho, P.L.; Tong, E.; Wang, L.; Xu, B. Nano Letters. 2003,3,1261-1263.
CrossRef - Schabes-Retchkiman, P.S.; Canizal, G.; Herrera-Becerra, R; Zorrilla, C.; Liu, H.B.; Ascencio, J.A. Opt Mater.2006, 29,95-99.
CrossRef - Gong, P. Nanotechnology. 2007,18,285-604.
CrossRef - Sheng, Y.; Narayanan, M.; Basha, S.; Elfasakhany, A.; Brindhadevi, K.; Xia, C. Pugazhendhi, A. Process Biochem. 2022,112,241–247.
CrossRef - Arivalagan, P.; Desika, P.; Jaya Mary, J.; Indira, K.; Rijuta Ganesh, S. Microb Pathogenesis. 2017,114,41–45.
- Anatol, D.; Bulcha, B.; Leta Tesfaye, J.; Boka, F.; Shanmugam, R.; Lalitha Priyanka, D. Nagaprasad, N.; Ramaswamy, K. J Nanomater. 2021,9,2021.
- Krishnaraj, C.; Jagan, E.G.; Rajasekar, S.; Selvakumar, P.; Kalaichelvan, P.T.; Mohan, N. Colloids Surf, B. 2010,76,50–56.
CrossRef - Panacek, A.; Kolar, M.; Vecerova, R.; Prucek, R.; Soukupova, J.; Krystof, V.; Hamal, P.; Zboril, R.; Kvítek, L. Biomat. 2009,30,6333–40.
CrossRef - Shrivastava, S.; Dash, D. J Nanotech.2009, 12,240–243.
CrossRef - Gurunathan, S.; Lee, K.; Kalishwaralal, K.; Sheikpranbabu, S.; Vaidyanathan, R.; Eom, S. Biomat. 2009,30,6341–50.
CrossRef - Rogers, J.V.; Parkinson, Y.W.; Choi, J.L.; Speshock, S.M.; Hussain, M.A. Nanoscale Res Lett. 2008, 3,129–133.
CrossRef - Nadworny, P.L.; Wang, J.; Tredget, E.E.; Burrell, R.E. Nanomed Nanotech Biol. Med. 2008,4,241–251.
CrossRef - Velusamy, P.; Das, J.; Pachaiappan, R.; Vaseeharan, B.; Pandian, K. Ind Crop Prod. 2015,66,103-109.
CrossRef - Halawani, E.M. J Biomater Nanobiotech. 2017,8,22-35.
- Mostafa, M.; Al-Emam, A.; Sayed, M.; Alamri, S.; Alghamdi, H.; Shati, A.; Alrumman, S.; Alfaify, A.K. Bangladesh J. Bot. 2020, 49, 1127-1134.
CrossRef - Saied, A.S.; Gebauer, J.; Hammer, K.; Buerkert, A. Resour. Crop Evol. 2008, 55,929-937.
CrossRef - Moustafa, M.F.; Hesham, A.L.; Quraishi, M.S.; Alrumman, S.A. Genet. Eng. Biotechnol. 2016,14,349-362.
CrossRef - Kumar, G.; Sangita, M.; Bappadity, M.; Kiran, M. J. of Drug Discovery & Herbal Res.2011,1,231-233.
- Nazif, N.M. Food Chem.2002,76,77–81.
CrossRef - Elgorban, A.M.; Al-Rahmah, A.N.; Sayed, S.R.; Hirad, A.; Mostafa, A.F.; Bahkali, A.H.; Biotechnol. Equip.2016,30, 299–304.
CrossRef - Gurunathan, S.; Kalishwaralal, K.; Vaidyanathan, R.; Venkataraman, D.; Pandian, S.K.; Muniyandi, J.; Hariharan, N.; Eom, S.H. Colloids and Surfaces B: Biointerfaces.2009,74, 328–335.
CrossRef - Zhang, X.; Liu, Z.; Shen, W.; Gurunathan, S. J. Mol. Sci.2016, 17,1534.
CrossRef - Abalkil, T.; Al harbi, S.A.; Salmen, S.H.; Wainwright, M. Biotechnol. Equip.2017,31,411–417.
CrossRef - Salmen, S.H.; Alwhibi, M.S.; Alharbi, SA. Ecol. Environ. Res. 2019,17,12869-12876.
- Salmen, S.H.; Alharbi, S.A. Green Chem. Lett. Rev.2020, 13, 1-5.
CrossRef - Karthik, C.; Suresh, S.; Sneha Mirulalini, G.; Kavitha, S.A. Inorganic and Nano-Metal Chem.2020, 50,606–612.
CrossRef - Dinesh, B.; Monisha, N.; Shalini, H.R.; Prathap, G.K.; Jagadeesh Poyya.; Manjula Shantaram.; Jayanth, S.; Hampapura.; Karigar, C.S.; Joshi, C.G.; SPECTROSCOPY LETTERS. 2022, 55, 20–34.
CrossRef - Lemike, E.E.; Fayemi, O.E.; Ekennia, A.C.; Onwudiwe, D.C.; Ebenso, E.E.; Molecules.2017,22, 701.
CrossRef - Sun, Q.; Cai, X.; Li, J.; Zheng, M.; Chen, Z.; Yu, C.P. Colloids Surf. A Physicochem. Eng. Aspects.2014, 444,226–231.
CrossRef - Carballo, T.; Gil, M.V.; Gómez, X.; González-Andrés, F.; Morán, A.; 2008, 19, 815–830.
CrossRef - Gomathi, M.; Rajkumar, P.V.; Prakasam, A.; Ravichandran, K. Resource Technology.2017, 3, 280–284.
CrossRef - Salmen, S.H. J. Chem.2020,36, 1189-1193.
CrossRef - Salmen, S.H.; Alkammash, N.M.; Alahmadi, T.A.; Alharbi, S.A. Chim. 2021,72,76-82.
CrossRef - Netala, V.R.; Kotakadi, V.S.; Bobbu, P.; Gaddam, S.A, Tartte, V. 3 Biotech. 2016, 6, 132–139.
CrossRef - Begum, N.A.; Mondal, S.; Basu, S.; Laskar, R.A.; Mandal, D. Colloids and Surfaces B: Biointerfaces. 2009,71, 113–118.
CrossRef - Song, J.Y.; Jang, H.K.; Kim, B.S. Process Biochemistry. 2009, 44, 1133–1138.
CrossRef - Morones, J.R.; Elechiguerra, J.L.; Camacho, A.; Holt, K.; Kouri, J.B.; Ramírez, J.T.; Yacaman, M.J. 2005, 16, 2346–2353.
CrossRef - Abul Qais, F.; Shafiq A Khan, H.M.; Husain, F.M.; Khan, R.A.; Alenazi, B.; Alsalme, A.; Ahmad, I. Chem. Appl. 2019, ID 4649506.
CrossRef
Accepted on: 28 Sep 2025

















