Based on Bioinformatics and Network Pharmacology Analyses, The Multi-System Mechanism by which Ge Gen Qin Lian Decoction Intervenes in Chemotherapy-induced Diarrhea through the Regulation of Key Genes and Signaling Pathways is Elucidated
1College of Pharmacy, Shaanxi University of Chinese Medicine, Xianyang, China
2Laboratory of Medicine, Datong Second People's Hospital (Datong Tumor Hospital), Datong, China
Corresponding Author E-mail: sxdteyyzx@163.com
DOI : http://dx.doi.org/10.13005/ojc/410504
ABSTRACT:Based on the well-documented efficacy of the classical formula GQD, traditionally used to “relieve exterior symptoms and clear internal heat, promoting diuresis,” this study investigates its mechanism in treating chemotherapy-induced diarrhea (CID) using network pharmacology and molecular docking. Chemical constituents of Pueraria lobata, Scutellaria baicalensis, Coptis chinensis, and Glycyrrhiza uralensis were retrieved from databases, and potential targets were predicted to construct a component–target database. CID-related targets were collected from Gene Cards, and common targets were analyzed through PPI networks using STRING. Core targets were further assessed via Bio GPS expression profiles. GEO datasets were analyzed to obtain differentially expressed genes, followed by GO and KEGG enrichment analyses to clarify biological processes and signaling pathways, while GSEA evaluated expression trends. Finally, molecular docking validated compound–target interactions. Network pharmacology identified 83 active ingredients and 37 potential targets, with epiberberine, coptisine, quercetin, beta-sitosterol, and formononetin as key components. GO and KEGG analyses revealed that GQD mainly modulates inflammatory responses and cancer-related pathways. The organ–target network indicated multi-component, multi-target, and multi-pathway effects. Docking results showed hydrogen bonding, hydrophobic, and other non-covalent interactions as crucial for compound–target binding. This study systematically elucidates the active components, potential targets, and molecular mechanisms of GQD against CID, providing a research strategy for understanding Chinese herbal formulas and supporting further development of GQD.
KEYWORDS:Chemotherapy-induced diarrhea; Gene Expression; GQD; Molecular Docking; Network Pharmacology
Introduction
Chemotherapy remains a cornerstone of cancer treatment; however, reducing its side effects is essential for improving patients’ quality of life.1 Chemotherapy-induced diarrhea (CID) is one of the most common adverse effects of antineoplastic drugs, and it accounts for more than 80% of chemotherapy-related side effects.2 Severe cases of CID (graded as 3 or 4) occur in over 30% of patients who are affected by this condition.3 Additionally, cancer itself has been reported to increase the risk of CID following radiation therapy, which may manifest as small bowel obstruction, abdominal pain, increased frequency of bowel movements, and loss of bowel control—symptoms that may persist for years or even a decade.4 CID primarily arises from the toxic effects of chemotherapeutic drugs on rapidly proliferating gastrointestinal epithelial cells, resulting in intestinal mucosal inflammation and disruption of the gut microbial balance. 5, Key mechanisms include apoptosis, upregulation of pro-inflammatory factors, and the spread of inflammation.6 These complex physiological processes pose significant challenges for the clinical management of CID.
Current therapeutic strategies for CID primarily involve pharmacological interventions, including the long-acting extended-release (LAR) formulation of octreotide7, 8 and loperamide.9 Octreotide (LAR) has shown efficacy as a second-line prophylactic treatment for CID,8 reducing its incidence and severity.10 However, its effectiveness can vary depending on patient characteristics, chemotherapy type, and study design, necessitating further systematic evaluation.7 Loperamide, an affordable and orally administered drug, is widely used for mild (grade 1 or 2) diarrhea.11 However, its efficacy in severe cases is limited,12, 13 and long-term use may be associated with a rare risk of serious cardiac events.14
From a TCM perspective, chemotherapy is considered a method of “attacking the evil,” which aligns with the TCM principles of “eliminating toxins” or “expelling pathogens”.15 However, while chemotherapy targets the disease, it also damages the body’s “positive qi,” particularly affecting spleen and stomach function, leading to deficiencies in qi, blood, yin, and bodily fluids.16 TCM integrates a holistic approach with modern scientific methodologies.17 utilizing both macro and micro diagnostic indicators to enhance the objectivity and applicability of treatment strategies.18 The combination of herbal medicine, acupuncture, moxibustion, and massage with chemotherapy has been shown to improve treatment efficacy.19 enhance immune function, reduce side effects, and restore the balance of yin and yang, thereby optimizing therapeutic outcomes and improving patients’ quality of life.20
Gegen-Qinlian Decoction (GQD), a classical formula from Zhang Zhongjing’s Treatise on Cold Damage Disorders, composed of Pueraria lobata, Scutellaria baicalensis, Coptis chinensis, and Glycyrrhiza uralensis. GQD exhibits anti-inflammatory, antioxidant, and antimicrobial properties and has demonstrated therapeutic efficacy in conditions such as type 2 diabetes mellitus (T2DM)21 and infectious lung injury.22 Moreover, GQD has been reported to enhance the efficacy of anticancer drugs while mitigating their adverse effects, particularly in alleviating CID.23 The mechanisms underlying GQD’s therapeutic effects involve multiple signaling pathways. Its flavonoids and alkaloids suppress the activation of NF-κB and MAPK signaling pathways, reducing inflammatory cytokine release and alleviating inflammation.24 Additionally, GQD modulates the Nrf2/HO-1 pathway to enhance cellular antioxidant capacity, mitigate oxidative stress, and protect the integrity of the epithelial barrier.25 Glycyrrhiza uralensis and its metabolites contribute to cell membrane repair, reinforce barrier function, and regulate chemotherapy-induced gut microbiota imbalances.26, 27 Furthermore, GQD may promote tissue repair and enhance drug tolerance via the PI3K-AKT signaling pathway.28
Due to its anti-inflammatory, antioxidant, barrier-protective, and microbiota-modulating effects, GQD has been suggested as a promising therapeutic option for CID. This study integrates network pharmacology, molecular docking, and multidimensional data analysis to systematically explore the therapeutic potential of GQD in CID treatment. By analyzing its core components and major active ingredients, we aim to elucidate its mechanisms of action and key therapeutic targets. This research not only clarifies the potential benefits of GQD for CID but also provides a scientific foundation for its clinical application, paving the way for future drug development targeting this condition. The research methodology is illustrated in Figure 1.
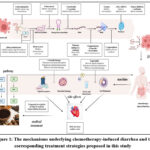 |
Figure 1: The mechanisms underlying chemotherapy-induced diarrhea and the corresponding treatment strategies proposed in this study |
Materials and Methods
Screening of targets for core drug active ingredients
Using the TCMSP Database and Analysis Platform29(https://www.tcmsp-e.com/), we retrieved the active components of the four herbs in Gegen Qinlian Decoction . The screening criteria were established as follows: molecular weight30(MW)≤500、oral bioavailability31(OB)≥30% and drug-like properties 32(DL)≥0.18. The identified active components were then imported into the “Swiss Target Prediction” database, where the biological species was specified as “Homo sapiens” and the prediction threshold was set to “Probability > 0”. This process yielded the target data associated with the active components.
Screening of disease targets associated with Chemotherapy-induced diarrhea
Using “chemotherapy-induced diarrhea” as the keyword, conduct a search for associated targets in the Gene Card33 database(https://www.genecards.org). Within this database, a higher score indicates a stronger correlation between the target and the condition. Finally, utilize a Venn diagram tool34 (https://bioinformatics.com.cn) to identify the common intersection of targets related to “disease-active ingredients”.35
Construction of a ‘core drug-active ingredient-CID disease target’ network
The “core drug–active compound–CID disease target” network was built using Cytoscape 3.10.0, with node color and size adjusted according to the Degree value.
Drawing of PPI networks
The overlapping targets between core drug targets and disease-related targets were uploaded to the STRING36 database (https://cn.stringdb.org/) to generate a PPI network. The species was specified as “Homo sapiens,” and the minimum required interaction score was set to ≥0.4. The obtained PPI network was subsequently imported into Cytoscape 3.10.0 for in-depth analysis. Key targets were identified by evaluating three topological metrics: DC, CC, and BC, which reflect the significance and connectivity of each node within the network. Furthermore, the MCODE plugin in Cytoscape 3.10.0 was applied to cluster and simplify the PPI network, using the following parameters: minimum MCODE score > 2, Node Score Cutoff = 0.2, k-core = 2, and maximum depth = 100.
Construction of organ-targets network
The treatment of diarrhea involves the coordinated function of multiple organs and tissues. However, the metabolic mechanisms of Ge Gen Qin Lian Decoction (GGQLD) in the body remain incompletely understood. To investigate its impact across multiple organs and tissues, we leveraged the Bio GPS37 database to retrieve mRNA expression data of the target genes in various human tissues and organ systems. The Bio GPS database provides comprehensive data on gene expression in normal human tissues, with a Z-value greater than 5 indicating significant gene expression in a specific organ or tissue. The organ-target network was constructed using Cytoscape 3.10.0 software.
GEO dataset gene expression differential analysis.
The GEO38 datasets(https://www.ncbi.nlm.nih.gov/)is a public repository for gene expression and high-throughput molecular data, encompassing clinical information from a wide range of diseases. Researchers can utilize this resource to retrieve mRNA expression profiles for various conditions.39 In the study of chemotherapy-induced diarrhea, we selected the GSE11223 dataset from the GEO database for analysis. This dataset comprises 202 samples, including 67 from patients with chemotherapy-induced diarrhea and 135 from healthy controls. DEGs between the patient and control groups were identified using the GEO2R tool, applying a threshold of log2 fold change > 1.2 and an adjusted p-value < 0.01. This analysis provides a foundation for elucidating the molecular mechanisms underlying chemotherapy-induced diarrhea.
GO enrichment assay and KEGG pathway analysis
The potential target sites of the core drugs were submitted to the Metscape40 platform (https://metascape.org/) with the organism set as “Homo sapiens”. The analysis was conducted with the following settings: a minimum overlap (Min Overlap) of 3, a P value cutoff of 0.01, and a minimum enrichment ratio (Min Enrichment) of 1.5. Following this, Gene Ontology (GO) and KEGG pathway enrichment analyses were carried out. The results were ranked based on log P values for enrichment, and the top 20 entries were selected. Bar charts were generated using relevant bioinformatics tools. The enriched pathways and targets were further analyzed by submitting them to the CNS Knowall platform (https://cnsknowall.com) to construct a “target-pathway” Sankey diagram.41 To clearly depict the interconnections among compounds, targets, and signaling pathways, a “compound–target–pathway” network was constructed using Cytoscape 3.10.0, followed by comprehensive analysis and assessment. Finally, cancer-related pathways were visualized using the KEGG PATHWAY42 database (https://www.kegg.jp).
Gene Set Enrichment Analysis (GSEA)
Enrichment analysis aids in identifying the molecular pathways potentially involved in GQD treatment of CID. Building on the results of KEGG enrichment analysis, GSEA analysis was employed to evaluate the expression patterns and overall trends of these pathways between the disease and control groups. In GSEA analysis,43 pathway enrichment significance is determined by an adjusted p-value < 0.05 and a q-value < 0.25, ensuring the statistical reliability and biological relevance of the findings. Through GSEA analysis, key signaling pathways that GQD may target can be thoroughly elucidated, offering new insights into the molecular mechanisms underlying its therapeutic effects on CID.
Gene survival analysis
In the GEPIA2 [44],we analyzed the overall survival (OS) of target genes across all cancer types. Based on the node degree values in the PPI network, the top two key genes were identified for further investigation. In the survival analysis, patients were divided into high-expression and low-expression groups, with each group representing half of the total cohort. The HR along with its 95% CI was computed to evaluate the association between gene expression levels and survival outcomes.
Molecular docking
The five most active compounds were obtained from the TCMSP database and converted into PDB format using Open Babel 3.1. The crystal structures of the two highest-ranking proteins were then retrieved from the RCSB Protein Data Bank [45] (https://www.rcsb.org). The protein structures were prepared using Auto Dock Tools 1.5.6 by adding hydrogen atoms, assigning partial atomic charges, and eliminating water molecules. The binding site for molecular docking was defined based on the location of the original ligand within the protein. After removing the original ligand, the prepared protein structure files were generated for subsequent docking studies. All active compounds were then docked into the predefined binding site using Auto Dock Tools 1.5.6. Finally, the docking results were visualized and analyzed using PyMOL 2.5 software.
Results
GQD active ingredients and targets
In the TCMSP database, a total of 129 active components and 275 targets were identified. The active components of Ge Gen Qin Lian Decoction are summarized in Table S1. As shown in Table S1, Pueraria lobata (GG) contributed 3 active components, Glycyrrhiza uralensis (GC) contributed 83, Scutellaria baicalensis (HQ) contributed 37, and Coptis chinensis (HL) contributed 13. Notably, β-sitosterol (MOL000358) is a common component shared between Pueraria lobata and Scutellaria baicalensis, while formononetin (MOL000392) is shared between Pueraria lobata and Glycyrrhiza uralensis. Additionally, berberine, epiberberine, and quercetin are common components found in Coptis chinensis, Scutellaria baicalensis, and Glycyrrhiza uralensis.
Disease targets
A total of 810 disease targets associated with chemotherapy-induced diarrhea were retrieved from the GeneCards database. The intersection of these targets with those corresponding to the core drugs identified in Section 2.1 was then determined. Ultimately, 36 common drug-disease targets were identified (see Figure 2), as detailed in Table 1.
Table 1: Targets shared by GQD and chemotherapeutic diarrhea
| No | Symbol | No | Symbol | No | Symbol |
| 1 | PTGER2 | 14 | CDC42 | 27 | TOP1 |
| 2 | IDO1 | 15 | EGFR | 28 | ABCB1 |
| 3 | HDAC9 | 16 | NOS2 | 29 | CNR1 |
| 4 | COMT | 17 | CA9 | 30 | BCHE |
| 5 | NR3C1 | 18 | TNF | 31 | LRRK2 |
| 6 | CNR2 | 19 | HTR3A | 32 | SIRT1 |
| 7 | PDE5A | 20 | MAOB | 33 | CES1 |
| 8 | AR | 21 | MIF | 34 | ABCG2 |
| 9 | PLG | 22 | PRKCA | 35 | CES2 |
| 10 | NR1H4 | 23 | CHRM3 | 36 | ACHE |
| 11 | ESR1 | 24 | XDH | 37 | DRD2 |
| 12 | PDE4A | 25 | ESR2 | ||
| 13 | PIK3CG | 26 | CYP19A1 |
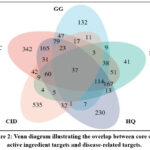 |
Figure 2: Venn diagram illustrating the overlap between core drug active ingredient targets and disease-related targets. |
Core Drug-Active Ingredient-CID Disease Target’ network
The “core drug–active ingredient–CID target” network was built using Cytoscape software (as shown in Figure 2), consisting of 311 nodes and 3,169 edges in total. The analysis revealed that the top five active ingredients ranked by node degree were berberine, coptisine, quercetin, β-sitosterol, and formononetin (see Table 2).
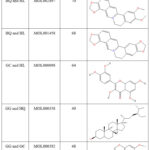 |
Table 2: GQD active ingredient information. |
Draw a PPI network diagram
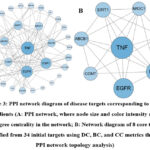
To explore the underlying mechanisms of Ge Gen Qin Lian Decoction in managing chemotherapy-induced diarrhea, a PPI network was established based on the identified target genes and analyzed using topological centrality metrics, including DC、BC and CC. According to the results presented in Figure 3A, applying a threshold of DC ≥ 1 yielded a network comprising 34 nodes and 109 edges. Further refinement with stringent criteria of DC ≥ 10, BC > 0.03, and CC > 0.54 resulted in a subnetwork of 8 key targets connected by 22 edges. The most significant targets identified were TNF, EGFR, ESR1, SIRT1, ABCB1, NR3C1, COMT, and MAOB. To visually represent the degree centrality of these pivotal targets, a horizontal bar chart was generated for the PPI network (Figure S2). Additionally, cluster analysis using the MCODE plugin in Cytoscape was conducted to identify highly interconnected subnetworks, which were subsequently categorized into two groups (Figure S3).
Construction and Analysis of the Organ-Target Network
Analysis of mRNA expression levels for the 37 targets within the PPI network, based on data from the BioGPS database, revealed that all 37 targets showed increased mRNA expression in tissues associated with diarrhea. Specifically, significant increases were observed in Liver (degree: 49), Whole Blood (degree: 36), CD14+ Monocytes (degree: 36), Small Intestine (degree: 35), Trigeminal Ganglion (degree: 34), and Colon (degree: 34). As illustrated in Figure 4, the tissue-target network comprises 51 nodes and 475 edges. The majority of these targets showed high expression levels in the tissues, indicating a strong association between these organs and chemotherapy-induced diarrhea targets. Further analysis revealed that the 14 involved tissues and organs are closely linked to immune function, blood circulation, and metabolism, suggesting that the therapeutic effects of Ge Gen Qin Lian Decoction may involve systemic organ systems.
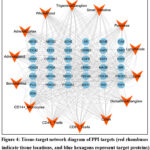 |
Figure 4: Tissue-target network diagram of PPI targets (red rhombuses indicate tissue locations, and blue hexagons represent target proteins). |
Differential gene expression analysis of the GEO database
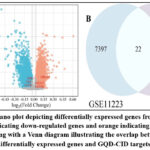
We obtained a gene dataset related to CID from the GEO database and conducted differential expression analysis using the GEO2R analysis tool, drawing a volcano plot of gene differential expression. In the volcano plot, the differential expression of the top-ranked genes in the PPI was displayed, among which TNF, EGFR, and ESR1 were all significantly downregulated. Furthermore, through the identification of differentially expressed genes using a significance threshold of p < 0.01 and intersecting them with the genes from the PPI network, a final set of 22 overlapping genes was obtained. These shared genes are likely to be critically involved in the initiation and progression of CID, offering valuable insights for uncovering its underlying molecular mechanisms.
To visually illustrate the results of differential gene expression (DGE) analysis, this study generated several visualization charts, including an adjusted p-value distribution plot (Adjusted P-value counts), a gene expression density plot (Expression density plot), a UMAP dimensionality reduction clustering plot (UMAP plot), and a moderated t-statistic plot (Moderated t statistic plot), as presented in Figure S4. These visualizations facilitate the evaluation of data quality, sample distribution, and the reliability of the analysis outcomes. (A) Adjusted p-value distribution plot: This chart evaluates the overall distribution of p-values from the differential expression analysis (DEA). A high concentration of genes with p-values in the P < 0.05 range suggests the presence of numerous significantly differentially expressed genes, reflecting strong statistical power. (B) Gene expression density plot: This plot assesses the effectiveness of data standardization. Ideally, the density curves of all samples should overlap extensively, indicating that technical variations have been effectively minimized and data comparability has been enhanced. (C) UMAP dimensionality reduction clustering plot: This visualization illustrates the overall relationships among samples. If the diseased and control groups form distinct clusters in the UMAP space, it indicates significant differences in gene expression patterns between the two groups, supporting the rationale for subsequent differential gene screening. (D) Moderated t-statistic plot: This chart evaluates the distribution characteristics of the moderated t-statistic calculated using the limma method. Ideally, data points should align closely with the theoretically predicted straight line, confirming that the statistical calculations in the differential expression analysis conform to theoretical expectations and enhancing result credibility. In conclusion, the findings in Figure S4 demonstrate effective data standardization, clear sample grouping, and sound statistical analysis, further reinforcing the accuracy and reliability of the research results.
GO enrichment analysis
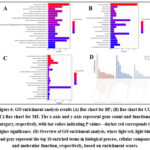
GO enrichment analysis was conducted through the Metscape database, and a total of 194 entries were screened out, including 124 BP, 24 CC, and 46 MF (see Figure S5).
To further explore the potential mechanisms of Ge Gen Qin Lian Decoction in treating chemotherapy-induced diarrhea, we conducted a visualization analysis based on the top 20 items ranked by P-values. The results covered three main categories: BP (Figure 6A), CC (Figure 6B), and MF (Figure 6C). Specifically, the key biological processes involved included inflammatory response and lipopolysaccharide response (Figure 6A); the primary cellular components affected were the plasma membrane, cell surface, and extrachromatin (Figure 6B); and the primary molecular functions identified included estrogen response element binding, nuclear receptor activity, and enzyme binding (Figure 6C). Additionally, the degree of enrichment was quantified through GO functional enrichment analysis, with enrichment scores presented in Figure 6D.
KEGG pathway analysis
KEGG pathway enrichment analysis was conducted on the 37 intersection genes, resulting in the identification of 28 pathways. The 20 most significantly enriched pathways were selected based on P-values and the number of associated genes (as shown in Figure 7). Subsequent analysis using KEGG bubble plots and pathway classification charts indicated that the core targets were predominantly linked to metabolic pathways, pathways related to cancer, and cytomegalovirus infection pathways. To more intuitively illustrate the relationships between components, targets, and pathways, a “component-target-pathway” network diagram was constructed (see Figure S6), comprising 97 nodes and 580 edges. Among these, the top five targets—PRKCA, EGFR, TNF, MAOB, and ESR2—were involved in 16, 11, 6, 6, and 5 pathways, respectively. Subsequently, a Sankey diagram was created using the CNS Knowall platform (refer to Figure S9). Finally, the cancer-related pathway was mapped within the KEGG PATHWAY database (see Figure S9), emphasizing key signaling pathways including PI3K-AKT, MAPK, and apoptosis.
 |
Figure 7: KEGG Enrichment Analysis Results (A) Bubble chart displaying the top 20 enriched pathways based on KEGG analysis; (B) Functional classification of the top 20 pathways. |
Enrichment analysis
In conjunction with KEGG enrichment analysis, this study focused on the alterations in the PI3K-AKT and MAPK signaling pathways, as illustrated in Figure 8. The enrichment analysis revealed that both pathways were significantly upregulated in the CID-related gene set, indicating their potential critical roles in the pathogenesis and progression of CID. Activation of the PI3K-AKT and MAPK pathways may promote cell proliferation, enhance inflammatory responses, and regulate the expression of pro-inflammatory factors, thereby exacerbating inflammatory damage. Furthermore, these pathways are critically involved in modulating immune responses and maintaining cell viability, which may impact intestinal barrier integrity and the secretion of inflammatory factors. Overall, these results shed new light on the molecular basis of CID and suggest promising avenues for developing targeted therapeutic interventions.
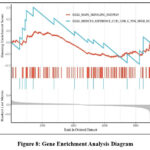 |
Figure 8: Gene Enrichment Analysis Diagram |
Gene Survival analysis
To investigate the impact of genes associated with chemotherapy-induced diarrhea on survival in various cancers, we conducted survival analysis using the GEPIA2 database for the EGFR (Figure S10A) and TNF (Figure S10B) genes. Patients were divided into high-expression and low-expression groups, each representing half of the cohort, and hazard ratios together with their corresponding 95% confidence intervals were computed. The results demonstrated that patients with lower expression levels of EGFR and TNF generally exhibited prolonged survival. These findings indicate that the two genes may be critically involved in cancer prognosis and hold promise as therapeutic targets for enhancing patient survival and reducing chemotherapy-associated diarrhea.
Molecular docking
To verify the targets identified via network pharmacology, molecular docking was conducted to evaluate the binding interactions between the selected active compounds and their corresponding targets. The top five targets and the top five compounds were selected for docking analysis based on their degree centrality rankings in the PPI network. The docking scores are presented in Figure S11. Notably, MOL000358 (β-sitosterol), MOL001458 (coptisine), and MOL002897 (berberine) exhibited relatively high binding affinities with both proteins, as detailed in Table 3.
Table 3: Docking Scores between TNF and EGFR and the Top 3 Active Ingredients in GQD
| NO | Target | MOL ID | Compounds | Docking Score |
| A | TNF | MOL000358 | beta-sitosterol | -9.08 kcal/mol |
| B | TNF | MOL001458 | coptisine | -7.39 kcal/mol |
| C | TNF | MOL002897 | epiberberine | -6.88 kcal/mol |
| D | EGFR | MOL000358 | beta-sitosterol | -8.89 kcal/mol |
| E | EGFR | MOL002897 | epiberberine | -7.08 kcal/mol |
| F | EGFR | MOL001458 | coptisine | -6.54 kcal/mol |
Analysis of the Combined Model
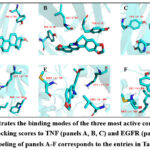
To investigate the binding modes and structural basis of interactions between active components and core targets, we utilized PyMOL for visualization analysis. Specifically, we examined the binding modes of the top three active ingredients listed in Table . The results revealed that beta-sitosterol exhibited the lowest binding energy when interacting with both TNF and EGFR targets, with values of -9.08 kcal/mol and -8.89 kcal/mol, respectively. In the TNF target, beta-sitosterol formed π-alkyl interactions with the key amino acid residue TYR151 [46]. Meanwhile, coptisine and epiberberine engaged in π-π stacking interactions with the critical residue TYR59 of the TNF target. The EGFR target interacted with beta-sitosterol via several non-covalent bonds, such as π-π stacking, π-alkyl, hydrophobic, and van der Waals interactions. Furthermore, coptisine and epiberberine established hydrogen bonds with the critical MET769 residue [47] of EGFR and simultaneously engaged in π-σ interactions with LEU694. These findings highlight the distinct binding patterns of different active components, particularly in their interactions with key amino acid residues (see Figure 9), providing a foundation for further research into the mechanisms of action and structural optimization of these compounds.
Discussion
Chemotherapy is a cornerstone of cancer treatment [48]. For instance, CPT-11, a camptothecin derivative and TOP1 inhibitor, forms CPT-11–TOP1–DNA complexes to block DNA replication, inducing cell cycle arrest and apoptosis [49]. Its metabolite SN-38 enhances anticancer effects by upregulating p53 [50] and pro-apoptotic proteins (e.g., Bax, caspase-3, caspase-9), while downregulating Bcl-xL [51]. Similarly, 5-FU metabolites inhibit thymidylate synthase, disrupt RNA/DNA synthesis, and induce apoptosis [52]. However, these agents damage intestinal epithelial cells, disrupting production-apoptosis balance, leading to barrier dysfunction and increased CID risk [53]. Thus, integrative strategies that target cancer and mitigate side effects are essential.
Despite the availability of clinical drugs for CID treatment, their efficacy remains limited, highlighting the need for more effective therapeutic strategies. Previous studies have explored approaches to mitigate chemotherapy-induced intestinal toxicity, including modulation of inflammatory bowel disease mechanisms [54]、 optimization of CPT-11 metabolism [55] and inhibition of oxidative stress [56]. However, these treatments are often administered only after the onset of diarrhea, forcing patients to interrupt chemotherapy, which not only compromises treatment efficacy but also significantly reduces their quality of life. Therefore, developing prophylactic or early-intervention therapies for CID has become an urgent clinical priority.
TCM offers a unique perspective on disease management, emphasizing individualized treatment based on the balance of qi, blood, yin, and yang. From a Western medicine standpoint, TCM compounds exert anticancer effects primarily by inhibiting tumor growth and metastasis while enhancing immune function [57, 58]. In addition to their potential in cancer therapy, herbal formulations serve as complementary and alternative medicines (CAM), alleviating chemotherapy-induced side effects and cancer-related fatigue (CRF) [59, 60]. Given the growing interest in integrative oncology, TCM-based interventions are increasingly recognized as promising therapeutic strategies [61].
Integrating TCM with modern oncology treatments has demonstrated benefits in symptom relief, improved quality of life, recurrence prevention, and prolonged survival [62]. TCM is rooted in a holistic approach, emphasizing individualized, evidence-based treatment, while Western medicine traditionally follows standardized treatment protocols. However, with the advent of personalized medicine, Western medical concepts are gradually aligning with TCM’s systemic regulatory principles [63]. Recent advances in evidence-based medicine have contributed to the standardization of TCM research methodologies, enhancing their scientific credibility and facilitating their integration into modern healthcare systems [64, 65]. Network pharmacology serves as a crucial bridge between TCM and modern evidence-based medicine. By employing systems biology approaches, it enables multi-level and multi-target analyses of TCM formulations, addressing limitations in traditional research methodologies and supporting the modernization of TCM [66]. Additionally, network pharmacology can integrate large-scale biological data, construct drug-target-disease interaction networks, and elucidate the multi-component, synergistic mechanisms of TCM formulas, thereby improving the reproducibility and scientific validation of TCM-based treatments. Clinically, it can optimize herbal formulations, predict drug-drug interactions, identify active ingredients with high efficacy and low toxicity, and guide personalized treatment strategies. Moreover, in drug discovery, network pharmacology accelerates active ingredient screening, target validation, and mechanism exploration, providing a scientific basis for innovative pharmaceutical development.
In this study, we constructed compound-target interaction networks using TCMSP, Gene Cards, and Swiss Target Prediction databases. Our results indicate that GQD exerts a multi-target, multi-pathway, and multi-organ regulatory effect in CID treatment by acting on key target genes such as TNF, EGFR, ESR1, SIRT1, and ABCB1. The primary mechanisms of GQD involve intestinal tissue repair, inhibition of intestinal epithelial apoptosis, and cancer cell apoptosis induction, contributing to CID relief. Based on PPI network analysis and a review of existing literature, GQD exerts anti-inflammatory effects by modulating the PI3K/AKT/NF-κB signaling cascade, leading to reduced production of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6, decreased neutrophil infiltration, and attenuation of oxidative stress. Moreover, GQD activates the Keap1/Nrf2 pathway, which strengthens the intestinal barrier by boosting antioxidant defenses and promoting the expression of tight junction proteins like ZO-1 and Occludin, thereby safeguarding intestinal epithelial cells and their mucosal secretions. [28]. Notably, TNF and its associated pro-inflammatory cytokines (e.g., IL-1β, COX-2, ICAM-1) play a pivotal role in CID pathogenesis. GQD’s ability to downregulate these inflammatory mediators suggests a protective effect against intestinal inflammation [67]. Furthermore, moderate modulation of EGFR and its downstream signaling pathways, including reduced PI3K and p-AKT expression, may help attenuate inflammation and further reinforce intestinal barrier function, contributing to CID symptom relief [68, 69].
Our findings also indicate that GQD modulates biological processes such as inflammatory response regulation, MAPK pathway activation, and intracellular signaling transduction, further supporting its role in CID management. Given that CID development is primarily driven by intestinal epithelial apoptosis and excessive inflammatory responses, targeting these pathways represents a key therapeutic strategy. KEGG pathway enrichment analysis suggests that in addition to its impact on cancer-related pathways, GQD may positively influence cancer prognosis while alleviating chemotherapy-induced intestinal inflammation.
While this study provides insights into GQD’s potential mechanisms in CID treatment through network pharmacology, it primarily relies on bioinformatics analyses. Although this study is exploratory, it provides a solid scientific foundation for future research on the application of GQD in managing CID. Nevertheless, further in vivo and in vitro experiments are required to directly validate the proposed active compounds, critical targets, and underlying molecular mechanisms.
Conclusion
Network pharmacology analysis reveals that Gegen-Qinlian Decoction (GQD) exerts a therapeutic effect on chemotherapy-induced diarrhea (CID) by targeting TNF, EGFR, ESR1, SIRT1, and ABCB1, and modulating key signaling pathways such as PI3K/AKT, NF-κB, MAPK, and TNF-related pathways. Through a multi-organ and multi-target regulatory mechanism, GQD contributes to intestinal barrier maintenance, inflammation suppression, and epithelial repair, aligning with TCM’s holistic view that emphasizes systemic regulation and overall balance.
However, this study is primarily based on bioinformatics approaches, necessitating further experimental validation in preclinical and clinical settings to confirm its efficacy and mechanism of action. Future research should focus on identifying GQD’s specific active components, validating their targets, and elucidating their action networks to strengthen the clinical applicability of this formula. The findings of this study provide a novel scientific perspective for advancing research and development of GQD as a potential treatment for chemotherapy-induced diarrhea.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Reference
- LIU Z, HUANG P, LAW S, et al. Preventive Effect of Curcumin Against Chemotherapy-Induced Side-Effects [J] Frontiers in pharmacology, 2018, 9: 1374.
CrossRef - STEIN A, VOIGT W, JORDAN K. Chemotherapy-induced diarrhea: pathophysiology, frequency and guideline-based management [J] Therapeutic advances in medical oncology, 2010, 2(1): 51-63.
CrossRef - MAROUN J A, ANTHONY L B, BLAIS N, et al. Prevention and management of chemotherapy-induced diarrhea in patients with colorectal cancer: a consensus statement by the Canadian Working Group on Chemotherapy-Induced Diarrhea [J] Current oncology (Toronto, Ont), 2007, 14(1): 13-20.
CrossRef - DENLINGER C S, BARSEVICK A M. The challenges of colorectal cancer survivorship [J] Journal of the National Comprehensive Cancer Network : JNCCN, 2009, 7(8): 883-93; quiz 94.
CrossRef - WADLER S, BENSON A B, 3RD, ENGELKING C, et al. Recommended guidelines for the treatment of chemotherapy-induced diarrhea [J] Journal of clinical oncology : official journal of the American Society of Clinical Oncology, 1998, 16(9): 3169-78.
CrossRef - MCQUADE R M, STOJANOVSKA V, ABALO R, et al. Chemotherapy-Induced Constipation and Diarrhea: Pathophysiology, Current and Emerging Treatments [J] Frontiers in pharmacology, 2016, 7: 414.
CrossRef - DENG C, DENG B, JIA L, et al. Efficacy of long-acting release octreotide for preventing chemotherapy-induced diarrhoea: protocol for a systematic review [J] BMJ open, 2017, 7(6): e014916.
CrossRef - ZIDAN J, HAIM N, BENY A, et al. Octreotide in the treatment of severe chemotherapy-induced diarrhea [J] Annals of oncology : official journal of the European Society for Medical Oncology, 2001, 12(2): 227-9.
CrossRef - GELLER R B, GILMORE C E, DIX S P, et al. Randomized trial of loperamide versus dose escalation of octreotide acetate for chemotherapy-induced diarrhea in bone marrow transplant and leukemia patients [J] American journal of hematology, 1995, 50(3): 167-72.
CrossRef - LI Y G, MENG X Y, YANG X, et al. Structural insights into somatostatin receptor 5 bound with cyclic peptides [J] Acta pharmacologica Sinica, 2024, 45(11): 2432-40.
CrossRef - SOMMER K, WULF S, GALLWAS J. [Measures for diarrhoea during chemotherapy and targeted therapy] [J] Deutsche medizinische Wochenschrift (1946), 2024, 149(10): 592-7.
CrossRef - LENFERS B H, LOEFFLER T M, DROEGE C M, et al. Substantial activity of budesonide in patients with irinotecan (CPT-11) and 5-fluorouracil induced diarrhea and failure of loperamide treatment [J] Annals of oncology : official journal of the European Society for Medical Oncology, 1999, 10(10): 1251-3.
CrossRef - BARBOUNIS V, KOUMAKIS G, VASSILOMANOLAKIS M, et al. Control of irinotecan-induced diarrhea by octreotide after loperamide failure [J] Supportive care in cancer : official journal of the Multinational Association of Supportive Care in Cancer, 2001, 9(4): 258-60.
CrossRef - DE LEMOS M L, GUENTER J, KLETAS V. Loperamide and cardiac events: Is high-dose use still safe for chemotherapy-induced diarrhea? [J] Journal of oncology pharmacy practice : official publication of the International Society of Oncology Pharmacy Practitioners, 2018, 24(8): 634-6.
CrossRef - ZHENG Z, CHO W C, XU L, et al. Lessons learnt from evidence-based approach of using chinese herbal medicines in liver cancer [J] Evidence-based complementary and alternative medicine : eCAM, 2013, 2013: 656351.
CrossRef - BAI B, MA Y, LIU D, et al. DNA damage caused by chemotherapy has duality, and traditional Chinese medicine may be a better choice to reduce its toxicity [J] Frontiers in pharmacology, 2024, 15: 1483160.
CrossRef - WANG S, LONG S, WU W. Application of Traditional Chinese Medicines as Personalized Therapy in Human Cancers [J] The American journal of Chinese medicine, 2018, 46(5): 953-70.
CrossRef - SO T H, CHAN S K, LEE V H, et al. Chinese Medicine in Cancer Treatment – How is it Practised in the East and the West? [J] Clinical oncology (Royal College of Radiologists (Great Britain)), 2019, 31(8): 578-88.
CrossRef - ZHANG Z, ZHU M J F I M. Holistic View of TCM on Cancer Integrative Therapy [J], 2023, 2(3): 159-67.
CrossRef - LIU S H, CHEN P S, HUANG C C, et al. Unlocking the Mystery of the Therapeutic Effects of Chinese Medicine on Cancer [J] Frontiers in pharmacology, 2020, 11: 601785.
CrossRef - XU Y, HUANG J, WANG N, et al. Network Pharmacology-Based Analysis and Experimental Exploration of Antidiabetic Mechanisms of Gegen Qinlian Decoction [J] Frontiers in pharmacology, 2021, 12: 649606.
CrossRef - LI W, DING Z, CHEN Y, et al. Integrated Pharmacology Reveals the Molecular Mechanism of Gegen Qinlian Decoction Against Lipopolysaccharide-induced Acute Lung Injury [J] Frontiers in pharmacology, 2022, 13: 854544.
CrossRef - LU J Z, YE D, MA B L. Constituents, Pharmacokinetics, and Pharmacology of Gegen-Qinlian Decoction [J] Frontiers in pharmacology, 2021, 12: 668418.
CrossRef - HU F Q, XIONG L Y, LI Z P, et al. Deciphering the shared mechanisms of Gegen Qinlian Decoction in treating type 2 diabetes and ulcerative colitis via bioinformatics and machine learning [J] Front Med, 2024, 11: 18.
CrossRef - ZHANG X Y, JI Z X, HE Q, et al. Gegen Qinlian Decoction inhibits liver ferroptosis in type 2 diabetes mellitus models by targeting Nrf2 [J] J Ethnopharmacol, 2025, 340: 15.
CrossRef - WANG B K, YE X L, ZHOU Y H, et al. Glycyrrhizin Attenuates <i>Salmonella</i> Typhimurium-Induced Tissue Injury, Inflammatory Response, and Intestinal Dysbiosis in C57BL/6 Mice [J] Frontiers in veterinary science, 2021, 8: 13.
CrossRef - LIU F, TANG X, MAO B Y, et al. Ethanol Extract of Licorice Alleviates HFD-Induced Liver Fat Accumulation in Association with Modulation of Gut Microbiota and Intestinal Metabolites in Obesity Mice [J] Nutrients, 2022, 14(19): 20.
CrossRef - CHEN J, LI M, CHEN R, et al. Gegen Qinlian standard decoction alleviated irinotecan-induced diarrhea via PI3K/AKT/NF-κB axis by network pharmacology prediction and experimental validation combination [J] Chinese medicine, 2023, 18(1): 46.
CrossRef - RU J, LI P, WANG J, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines [J] Journal of cheminformatics, 2014, 6: 13.
CrossRef - XIAO Y G, WU H B, CHEN J S, et al. Exploring the Potential Antidepressant Mechanisms of Pinellia by Using the Network Pharmacology and Molecular Docking [J] Metab Brain Dis, 2022, 37(4): 1071-94.
CrossRef - XU X, ZHANG W, HUANG C, et al. A novel chemometric method for the prediction of human oral bioavailability [J] International journal of molecular sciences, 2012, 13(6): 6964-82.
CrossRef - ZHANG W D, CHEN Y, JIANG H H, et al. Integrated strategy for accurately screening biomarkers based on metabolomics coupled with network pharmacology [J] Talanta, 2020, 211: 13.
CrossRef - STELZER G, DALAH I, STEIN T I, et al. In-silico human genomics with GeneCards [J] Human genomics, 2011, 5(6): 709-17.
CrossRef - LAM F, LALANSINGH C M, BABARAN H E, et al. VennDiagramWeb: a web application for the generation of highly customizable Venn and Euler diagrams [J] BMC bioinformatics, 2016, 17(1): 401.
CrossRef - WANG X, WU L, YU M, et al. Exploring the molecular mechanism of Epimedium for the treatment of ankylosing spondylitis based on network pharmacology, molecular docking, and molecular dynamics simulations [J] Molecular diversity, 2024.
CrossRef - SZKLARCZYK D, KIRSCH R, KOUTROULI M, et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest [J] Nucleic acids research, 2023, 51(D1): D638-d46.
CrossRef - WU C, JIN X, TSUENG G, et al. BioGPS: building your own mash-up of gene annotations and expression profiles [J] Nucleic acids research, 2016, 44(D1): D313-6.
CrossRef - BARRETT T, WILHITE S E, LEDOUX P, et al. NCBI GEO: archive for functional genomics data sets–update [J] Nucleic acids research, 2013, 41(Database issue): D991-5.
CrossRef - YANG J, ZHUANG Q, TANG K, et al. Exploring the action mechanism of Gegensan in the treatment of alcoholic liver disease based on network pharmacology and bioinformatics [J] Medicine, 2024, 103(25): e38315.
CrossRef - ZHOU Y, ZHOU B, PACHE L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets [J] Nature communications, 2019, 10(1): 1523.
CrossRef - SOUNDARARAJAN K, HO H K, SU B. Sankey diagram framework for energy and exergy flows [J] Appl Energy, 2014, 136: 1035-42.
CrossRef - KANEHISA M, FURUMICHI M, SATO Y, et al. KEGG: biological systems database as a model of the real world [J] Nucleic acids research, 2024.
CrossRef - SUBRAMANIAN A, TAMAYO P, MOOTHA V K, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles [J] Proc Natl Acad Sci U S A, 2005, 102(43): 15545-50.
CrossRef - WU C, MACLEOD I, SU A I. BioGPS and MyGene.info: organizing online, gene-centric information [J] Nucleic acids research, 2013, 41(Database issue): D561-5.
CrossRef - GOODSELL D S, ZARDECKI C, DI COSTANZO L, et al. RCSB Protein Data Bank: Enabling biomedical research and drug discovery [J] Protein science : a publication of the Protein Society, 2020, 29(1): 52-65.
CrossRef - HE M M, SMITH A S, OSLOB J D, et al. Small-molecule inhibition of TNF-alpha [J] Science (New York, NY), 2005, 310(5750): 1022-5.
CrossRef - PARK J H, LIU Y, LEMMON M A, et al. Erlotinib binds both inactive and active conformations of the EGFR tyrosine kinase domain [J] The Biochemical journal, 2012, 448(3): 417-23.
CrossRef - GILAD Y, GELLERMAN G, LONARD D M, et al. Drug Combination in Cancer Treatment-From Cocktails to Conjugated Combinations [J] Cancers, 2021, 13(4).
CrossRef - BAILLY C. Irinotecan: 25 years of cancer treatment [J] Pharmacological research, 2019, 148: 104398.
CrossRef - TAKEBA Y, KUMAI T, MATSUMOTO N, et al. Irinotecan activates p53 with its active metabolite, resulting in human hepatocellular carcinoma apoptosis [J] Journal of pharmacological sciences, 2007, 104(3): 232-42.
CrossRef - RUDOLF E, KRALOVA V, RUDOLF K, et al. The role of p38 in irinotecan-induced DNA damage and apoptosis of colon cancer cells [J] Mutation research, 2013, 741-742: 27-34.
CrossRef - LONGLEY D B, HARKIN D P, JOHNSTON P G. 5-fluorouracil: mechanisms of action and clinical strategies [J] Nature reviews Cancer, 2003, 3(5): 330-8.
CrossRef - XIANG J, WANG H, TAO Q, et al. CDK4/6 inhibitor modulating active and quiescent intestinal stem cells for prevention of chemotherapy-induced diarrhea [J] The Journal of pathology, 2023, 260(3): 235-47.
CrossRef - PAPADAKIS K A. Chemokines in inflammatory bowel disease [J] Curr Allergy Asthma Rep, 2004, 4(1): 83-9.
CrossRef - MATHIJSSEN R H J, VAN ALPHEN R J, VERWEIJ J, et al. Clinical pharmacokinetics and metabolism of irinotecan (CPT-11) [J] Clin Cancer Res, 2001, 7(8): 2182-94.
- BORDINI H P, KREMER J L, FAGUNDES T R, et al. Protective Effect of Metformin in an Aberrant Crypt Foci Model Induced by 1,2-Dimethylhydrazine: Modulation of Oxidative Stress and Inflammatory Process [J] Mol Carcinog, 2017, 56(3): 913-22.
CrossRef - YU Y X, WANG S, LIU Z N, et al. Traditional Chinese medicine in the era of immune checkpoint inhibitor: theory, development, and future directions [J] Chinese medicine, 2023, 18(1): 59.
CrossRef - CHEN F, LI J, WANG H, et al. Anti-Tumor Effects of Chinese Medicine Compounds by Regulating Immune Cells in Microenvironment [J] Frontiers in oncology, 2021, 11: 746917.
CrossRef - HUEBNER J, MUENSTEDT K, MUECKE R, et al. Is there level I evidence for complementary and alternative medicine (CAM) in oncology? An analysis of Cochrane Reviews [J] Trace Elem Electrolytes, 2013, 30(1): 29-34.
CrossRef - LIN J T, YANG T, CHEN W M, et al. Zhengyuan capsules for the treatment of chemotherapy-induced cancer-related fatigue in stage IIIB-IV unresectable NSCLC: study protocol for a randomized, multi-center, double-blind, placebo-controlled clinical trial [J] J Thorac Dis, 2022, 14(11): 4560-70.
CrossRef - XIA C, DONG X, LI H, et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants [J] Chinese medical journal, 2022, 135(5): 584-90.
CrossRef - JIAO L, BI L, LU Y, et al. Cancer chemoprevention and therapy using chinese herbal medicine [J] Biological procedures online, 2018, 20: 1.
CrossRef - LV W. Understanding traditional Chinese medicine [J] Hepatobiliary surgery and nutrition, 2021, 10(6): 846-8.
CrossRef - TIAN J J, LEVY M, SINNOTT R. A new paradigm of studying Traditional Chinese Medicine with evidence-based research [J] Pharmacological research, 2021, 172: 105858.
CrossRef - WU P, LI J, YAN H X, et al. Status and prospect of international standardization of TCM diagnosis [J] Pharmacological research, 2021, 171: 105746.
CrossRef - WANG Q, WANG X, WU K, et al. Network Pharmacology Unveils Multi-Systemic Intervention of Panax notoginseng in Osteoporosis via Key Genes and Signaling Pathways [J] Endocrine, metabolic & immune disorders drug targets, 2025.
CrossRef - WU Y, WANG D, YANG X, et al. Traditional Chinese medicine Gegen Qinlian decoction ameliorates irinotecan chemotherapy-induced gut toxicity in mice [J] Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie, 2019, 109: 2252-61.
CrossRef - WEI M, LI H, LI Q, et al. Based on Network Pharmacology to Explore the Molecular Targets and Mechanisms of Gegen Qinlian Decoction for the Treatment of Ulcerative Colitis [J] BioMed research international, 2020, 2020: 5217405.
CrossRef - LIU X, FAN Y, DU L, et al. In Silico and In Vivo Studies on the Mechanisms of Chinese Medicine Formula (Gegen Qinlian Decoction) in the Treatment of Ulcerative Colitis [J] Frontiers in pharmacology, 2021, 12: 665102.
CrossRef
Accepted on: 27 Sep 2025
Second Review by: Dr. Arvind Desai
Final Approval by: Dr. Ayssar Nahle