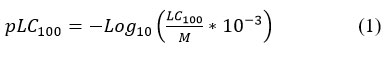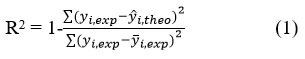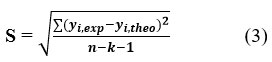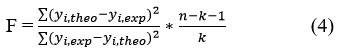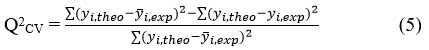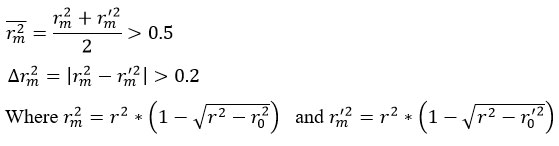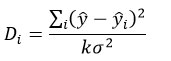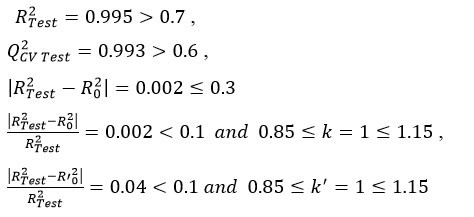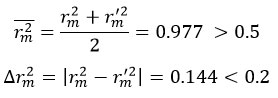Thiazolidinone Derivatives as Next-Generation Antifungal Scaffolds: Synthesis and QSAR Study
1Laboratoire de Constitution et Réaction de la Matière, UFR Sciences des Structures de la Matière et Technologie, Université Félix Houphouët-Boigny, Abidjan, Côte d’Ivoire.
2Département de Chimie Thérapeutique et Chimie Organique, UFR Sciences Pharmaceutique et Biologique, Université Félix Houphouët-Boigny, Abidjan, Côte d’Ivoire.
3Département des Sciences et de Technologie Agro-industrielles (STAgI), UFR Agriculture, ressources halieutiques et agro-industrie (ARHAI), Université de San Pedro, San Pedro, Côte d’Ivoire.
Corresponding author E-mail: souleydestras@yahoo.fr and tchambaga01@yahoo.fr
DOI : http://dx.doi.org/10.13005/ojc/410525
ABSTRACT:A novel series of 1,3-thiazolidin-4-one-5-arylidene derivatives was synthesized and evaluated for antifungal activity against Aspergillus flavus. The chemical structures were characterized by ¹H/¹³C NMR spectroscopy, along with HRMS. In vitro assays revealed moderate to strong activity, with MICs ranging from 15.565 to 238.942 µM, compared with itraconazole as a reference. Structure–activity analysis indicated that para-substituted electron-donating groups enhanced biological affinity. Compound 7b (4-dimethylamine) exhibited the highest potency. A QSAR study on derivatives 7a-j identified dipole moment (μD), lipophilicity (LogP), and polarizability (Pol) as key descriptors, with lipophilicity emerging as the most influential. Lower polarity favored stronger antifungal activity, and all derivatives except 7a contributed significantly to the predictive model. These findings highlight the promise of thiazolidinone scaffolds as viable leads for next-generation antifungal agents.
KEYWORDS:Antifungal activity; Aspergillus flavus; QSAR study, Structure–activity relationship; Thiazolidinone derivatives
Introduction
Fungal infections remain a serious global health concern, particularly in immunocompromised patients. Species of the genus Aspergillus are widespread saprophytic molds, typically harmless in the environment,1 but capable of causing life-threatening infections under specific conditions. Aspergillus fumigatus, whose airborne spores (conidia) humans inhale daily, is of particular concern2,3. While healthy individuals can eliminate these spores through pulmonary defenses, immunocompromised patients,4 such as those undergoing chemotherapy, corticosteroid treatment, or living with chronic diseases like diabetes or HIV/AIDS, are highly susceptible to invasive pulmonary mycoses.5 The clinical management of aspergillosis is becoming increasingly difficult due to the growing resistance of Aspergillus strains. Widespread use of azole fungicides in agriculture has driven selective pressure, leading to the emergence of resistant genotypes with cross-resistance to multiple antifungal classes. Alarmingly, some isolates show a predisposition to acquire resistance even to next-generation antifungal agents, partly due to genetic variants affecting DNA mismatch repair.6 Current therapies rely on the main classes of antifungals, such as allylamines, azoles, and pyrimidine analogues. However, these agents are limited by rising drug resistance,7 high costs, toxicity, and poor efficacy against biofilm-associated infections,8,9 which are notoriously difficult to eradicate.10 This situation poses a pressing public health challenge, with the looming risk of a future in which effective antifungal therapies may be unavailable.
To overcome these challenges, the development of new antifungal scaffolds is urgently required. Current approaches focus on designing combination therapies and novel molecules capable of inhibiting both fungal proliferation and biofilm formation.11 Among the various chemical classes under investigation, thiazolidinones become particularly popular. These heterocyclic compounds display a broad spectrum of biological activities, notably anti-inflammatory effects,12–14 antibacterial,15,16 antifungal,17,18 antimalarial,19 antimicrobial,20,21 antitumor22,23 and antidiabetic activity.24,25 Chemical modifications of the thiazolidin-4-one framework have generated derivatives with remarkable therapeutic promise, positioning this scaffold as a compelling platform for antifungal drug development.
This study explores a novel series of 5-arylidene-2-phenyl-3-(phenylamino)thiazolidin-4-one derivatives for antifungal activity against clinical strains of Aspergillus flavus. Alongside biological testing, a Quantitative Structure–Activity Relationship (QSAR) analysis was conducted to identify key molecular descriptors associated with antifungal potency. Overall, this work seeks to advance the development of safer and more effective antifungal agents capable of addressing the shortcomings of current therapies.
Materials and Methods
Materials
Chemistry Material
All reagents were commercially obtained (Aldrich, Fischer Scientific, France) and used as received. Reactions were monitored by TLC on Merck silica gel plates and visualized under UV light (254/365 nm). Products were purified via silica gel column chromatography, and melting points were measured using a Kofler bench (40–265 °C). Structures were confirmed by 1H and 13C NMR spectroscopy (Bruker 300/400 MHz, TMS as internal standard), with standard multiplicity notations (s, d, dd, t, q, m). Mass spectra were recorded in ESI mode on a JEOL JMS DX300 instrument. All chemical structures were drawn using ChemDraw (PerkinElmer), as presented in Figures 1–3.
Biological Materials
The synthesized compounds were assessed for antifungal activity against a clinical strain of Aspergillus flavus obtained from the Angré University Hospital Center in Côte d’Ivoire. Antifungal susceptibility was first assessed using the agar diffusion method.26 The test set consisted of ten synthesized derivatives, namely 1,3-thiazolidin-4-one-5-arylidene compounds 7a–j.
QSAR Materials
A QSAR study was conducted on compounds (7a–j). Seven molecules (7a, 7c, 7d, 7e, 7f, 7h, and 7i) were used as the training set, while three molecules (7b, 7g, and 7j) were reserved for external validation. The antifungal activities were determined across a concentration range of 12.5–102 μg/mL. This range enabled the establishment of quantitative correlations between antifungal potency and selected molecular descriptors. To allow numerical comparison, biological activity was expressed as the negative logarithm of the minimum inhibitory concentration -Log10(MIC) providing higher numerical values for more active compounds.27,28 Antifungal efficacy was further expressed using the parameter pLC₁₀₀, defined by the following equation (1):
where M represents the molecular weight (g/mol) and LC₁₀₀ is the lethal concentration required to eliminate 100% of the Aspergillus flavus population under the test conditions (µg/mL).
Methods
Chemistry Method
General procedure for the synthesis of 5-arylidene-2-phenyl-3-(phenylamino)thiazolidin-4-one 7 a-j
Compound 5 (1.11 mmol, 1.0 equiv) was dissolved in ethanol (2 mL) with sodium ethanolate (2.78 mmol, 2.5 eq.), and stirred at room temperature for 30 min. Benzaldehyde or substituted derivatives 6 (1.66 mmol, 1.5 equiv) were then added, and the reaction mixture was heated at 70 °C until completion, monitored by TLC. After cooling, the mixture was neutralized with 10% aqueous acetic acid, and the precipitate obtained was filtered, dried, and recrystallized from ethanol–water (80:20) to afford the target compounds 7a–j in 50–86% yield.
5-Benzylidene-2-phenyl-3-(phenylamino)thiazolidin-4-one 7a
Brown crystal, yield 70%, m.p.192-194°C, IR (ATR) (cm-1): 3257.6 ; 3098.1 ; 1612.0; 1603.2 ; 1580.0; 1036.7; 1042.6; 962.7. 1H NMR (400 MHz, CDCl3) δ: 7.91-7.82 (m, 2H), 7.83 (s, 1H), 7.46–6.83 (m, 13H), 5.92 (s, 1H), 5.76 (br, 1H). 13C NMR (101 MHz, CDCl3) δ : 164.4, 145.6, 136.8, 134.3, 133.4, 129.8, 129.1, 129.0, 128.8, 128.4, 128.2, 128.1, 126.0, 120.1, 112.8, 65.8. HRMS (ESI) Calc. for C22H18N2OS (M + H+) = 359.1140 Found = 359.1143.
5-(4-(Dimethylamino)benzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4- one 7b
Brown crystal, yield 50%, m.p. 136-138°C, IR (ATR) (cm-1): 3257.0; 1726.0, 1461.7, 1247–1192, 1050 ; 1H NMR (400 MHz, CDCl3) δ: 8.03 (s, 1H), 7.46–6.83 (m, 14H) 5.76 (br, 1H), 5.67 (s, 1H), 3.02 (s, 6H). 13C NMR (101 MHz, CDCl3) δ : 165.5, 144.7, 137.3, 135.3, 134.4, 129.7, 129.3, 129.1, 129.0, 128.6, 128.4, 128.3, 126.2, 120.1, 112.8, 64.9, 29.7. HRMS (ESI) Calc. for C24H23N3OS (M + H+) = 402.1562 Found = 402.1359.
5-(4-Hydroxybenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7c
Yellow crystal, yield 65%, m.p. 160-162°C, IR (ATR) (cm-1): 3520.3 , 3325.4; 1735.4, 1458.3 1H NMR (400 MHz, CDCl3) δ: 7.80-7.71 (m, 2H), 7.66 (s, 1H), 7.36–6.53(m, 14H), 5.90(s, 1H), 5.74(br, 1H). 13C NMR (101 MHz, CDCl3) δ : 164.4, 145.5, 136.7, 134.3, 133.4, 129.5, 129.1, 129.0, 128.7, 128.4, 128.2, 128.1, 125.9, 120.0, 112.7, 65.7. HRMS (ESI) Calc. for C22H18N2O2S (M + H+) = 375.1089 Found = 375.1102.
5-(4-Nitrobenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7d
Brown crystal, yield 86%, m.p. 130-132°C, IR (ATR) (cm-1): 3250.4 ; 3059.7 ; 1675.0; 1600.9 ; 1569.8; 1460.5-1582.1 ; 1104.8; 1046.2; 965.0. 1H NMR (300 MHz, CDCl3) δ 7.60 (s, 1H), 7.57–7.40 (m, 2H), 7.38–7.25 (m, 4H), 7.20–7.10 (m, 4H), 7.15–6.91 (m, 2H), 6.79–6.69 (m, 2H), 5.90 (s, 1H), 5.85 (br, 1H). 13C NMR (75 MHz, CDCl3) δ: 164.93, 137.1, 134.8, 129.7, 129.4, 129.2, 128.8, 128.6, 127.5, 127.4, 122.2, 122.0, 114.2, 65.1. HRMS (ESI) Calc. for C22H17N3O3S (M + Na+) = 426.0991 Found = 426.0992.
5-(4-Fluorobenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7e
Yellow crystal, yield 66%, m.p. 160-162°C, IR (ATR) (cm-1): 3257.6; 3061.0; 1678.7 ; 1600.9 ; 1460.5-1582.1; 1302.4; 1105.2; 1047.3; 960.5. 1H NMR (300 MHz, CDCl3) δ (ppm) 7.67 (s, 1H), 7.61–7.44 (m, 3H), 7.44–7.31 (m, 4H), 7.31–7.06 (m, 4H), 7.03–6.69 (m, 3H), 5.94 (s, 1H), 5.89 (br, 1H). 13C NMR (75 MHz, CDCl3) δ: 163.8, 136.1, 134.0, 129.6, 129.3, 129.1, 128.6, 128.4, 127.5, 127.3, 122.1, 122.0, 114.1, 61.4. HRMS (ESI) Calc. for C22H17FN2OS (M + Na+) = 399.1046 Found = 399.1044.
5-(4-Methoxybenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7f
Yellow crystal, yield 68%, m.p. 146-148°C, IR (ATR) (cm-1): 3257.77 ; 3000.0 ; 1678.07; 1598.9; 1492.9-1568.1; 1342.4 -1379.1; 1253.7; 1099.4;1029.9; 966.3; 833.2; 700.0 -752.2 . 1H NMR (400 MHz, CDCl3) δ :7.73-6.64 (m, 15H), 5.92 (s, 1H), 3.84 (s, 3H). 13C NMR (101 MHz, CDCl3) δ :165.2, 159.9, 145.0, 137.3, 135.3, 129.7, 129.4, 129.3, 129.2, 128.6, 128.4, 127.6, 127.5, 127.3, 126.2, 122.0, 120.1, 114.3, 114.1, 112.8, 61.54, 55.4. HRMS (ESI) Calc. for C23H20N2O2S (M + H+) = 389.1245 Found = 389.1248
5-(2,4-Dichlorobenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7g
Brown crystal, yield 80%, m.p. 72–73 °C, IR (ATR) (cm-1): 3257.7; 3061.0; 1687.7; 1600.9; 1465.9-1581.6; 1384.8; 1105.2; 1047.3; 960.5; 854.4; 748.3; 651.9-698.0 . 1H NMR (400 MHz, CDCl3) δ :7.95 (s, 1H), 7.48 (d, J = 2.2 Hz, 1H), 7.45–7.33 (m, 4H), 7.37–7.25 (m, 3H), 7.24 (dd, J = 6.9, 1.5 Hz, 2H), 6.96 (t, J = 7.4 Hz, 1H), 6.77–6.69 (m, 2H), 5.93 (s, 1H, CH), 5.87 (br, 1H). 13C NMR (101 MHz, CDCl3) δ :164.2, 144.7, 136.6, 135.7, 134.6, 131.5, 130.0, 129.9, 129.5, 129.4, 129.33, 127.4, 127.1, 122.2, 114.3, 61.7. HRMS (ESI) Calc. for C22H16Cl2N2OS (M + Na+) = 449.0360 Found = 449.0363.
5-(2,4-Dimethoxybenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4- one 7h
Yellow crystal, yield 70%, m.p. 164–166 °C, IR (ATR) (cm-1): 3246.2; 3018.6; 2835.36-2962.6; 1674.2; 1597.06; 1458.1- 1573.9; 1334.74-1392.6; 1257.5; 1099.4;1035.7; 960.5; 796.6-885.3; 700.0-748.3. 1H NMR (400 MHz, CDCl3) δ :7.43–7.28 (m, 2H), 7.39 (s, 1H), 7.32–7.19 (m, 6H), 6.94 (q, J = 6.7 Hz, 2H), 6.73 (t, J = 8.8 Hz, 2H), 6.57–6.43 (m, 1H), 5.88 (s, 1H), 5.77 (br, 1H), 3.85 (s, 6H). 13C NMR (101 MHz, CDCl3) δ :169.6, 165.4, 161.6, 159.3, 144.7, 130.0, 129.5, 129.3, 129.1, 127.4, 122.3, 122.0, 121.8, 114.2, 114.0, 104.6, 98.5, 62.0, 55.5, 55.4. HRMS (ESI) Calc. for C24H22N2O3S (M + Na+) = 441.1351 Found = 441.1351.
5-(3-Bromobenzylidene)-2-phenyl-3-(phenylamino)thiazolidin-4-one 7i
Brown crystal, yield 75%, m.p. 73–76 °C, IR (ATR) (cm-1): 3403.4; 3060.8; 1685.1; 1600.0; 1465.2-1580.2; 1380.7; 1105.2; 1047.3; 965.2; 625.0 .1H NMR (300 MHz, CDCl3) δ :7.45 (s, 1H), 7.40–7.24 (m, 8H), 7.22–7.06 (m, 4H), 7.04–6.67 (m, 2H), 5.90 (s, 1H), 5.87 (br, 1H). 13C NMR (75 MHz, CDCl3) δ :165.8, 136.4, 133.8, 129.4, 129.1, 129.0, 128.3, 128.2, 127.3, 127.2, 122.2, 121.7, 114.0, 61.2. HRMS (ESI) Calc. for C22H17BrN2OS (M + H+) = 436.0245 Found = 436.0242.
2-Phenyl-3-(phenylamino)-5-(thiophen-2-ylmethylene)thiazolidin-4-one 7j
Brown crystal, yield 80%, m.p. 226-228°C, IR (ATR) (cm-1): 3244.2; 3016.4-3103.4; 2881.6-2999.3; 1676.1; 1598.9; 1492.9 ; 1444.6 ; 1103.2; 1024.2; 966.6; 680.9- 752.2 . 1H NMR (400 MHz, CDCl3) δ :7.93 (s, 1H), 7.54 (d, J = 5.1 Hz, 1H), 7.46–7.32 (m, 6H), 7.26 (d, J = 7.8 Hz, 2H), 7.21–6.75 (m, 4H), 5.98 (br, 1H), 5.87 (s, 1H). 13C NMR (101 MHz, CDCl3) δ:164.7, 144.8, 139.2, 137.0, 130.2, 129.8, 129.4, 129.27, 128.7, 128.0, 127.5, 122.0, 120.6, 120.3, 114.1, 62.0. HRMS (ESI) Calc. for C20H16N2OS2 (M + Na+) = 387.0704 Found = 387.0707.
Biological Method
The synthesized compounds (e.g, compound 7a illustrated in Figure 1) were dissolved using dimethyl sulfoxide (DMSO) and distilled water as solvents. Antifungal activity was evaluated against Aspergillus flavus and compared with the reference drug Itraconazole. The fungal strain was supplied and identified by the Parasitology-Mycology Unit, Department of Medical Biology, Angré University Hospital. The antifungal activity was determined using the broth microdilution method in 96-well microplates, designed to establish the Minimum Inhibitory Concentration (MIC) of each compound. This assay involved exposing an inoculum of A. flavus to serial dilutions of the test compounds. Fungal growth inhibition was assessed by monitoring mitochondrial dehydrogenase activity through colorimetric detection with Methyl Thiazolyl Tetrazolium (MTT) chloride. Active cells convert the yellow MTT solution into purple formazan, whereas the absence of color change indicates inhibition of growth. The MIC was defined as the lowest concentration preventing purple coloration.29-31 The fungal inoculum was prepared from 7-day-old Aspergillus fumigatus colonies grown on Sabouraud chloramphenicol agar. Conidia were harvested in 5 mL of tryptone soy broth (OXOID®) and homogenized by vortexing for 15-20 seconds. Spore concentration was determined by Malassez cell counting and adjusted to 0.5 McFarland turbidity (≈0.4–5 × 10⁶ spores/mL). A 50-fold dilution yielded the working suspension, corresponding to approximately 0.08–1 × 10⁵ CFU/mL.32 Stock solutions of the compounds (1 mg/mL) were prepared in DMSO or water. Serial two-fold dilutions were performed in microplates containing tryptone soy broth, followed by the addition of the fungal inoculum. Plates were incubated at 37 °C for 7 days, and growth was assessed using MTT (2.5 mg/mL). Experiments were performed in duplicate and repeated twice for reproducibility.
QSAR Method
Quantum chemical calculations were carried out with Gaussian 09 using DFT to obtain molecular descriptors for QSAR analysis.33-37 Except for lipophilicity, calculated with A/logPS, descriptors were derived from geometry optimization and vibrational frequency calculations at the B3LYP/6-31+G(d,p) level. QSAR models were developed using multiple linear regression and validated with Microsoft Excel38 and validated using XLSTAT.39
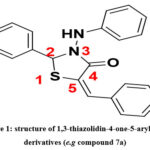 |
Figure 1: structure of 1,3-thiazolidin-4-one-5-arylidene derivatives (e.g compound 7a). |
Molecular descriptors
Twelve molecular descriptors were calculated for QSAR modeling, including energies (HOMO, LUMO, ΔE), chemical reactivity parameters, dipole moment, lipophilicity (LogP), and polarizability. The most predictive model combined LogP, dipole moment, and polarizability, with lipophilicity highlighted as a key factor for drug-likeness according to Lipinski’s rule. 40 The dipole moment , reflects the strength of intermolecular interactions,41 while polarizability is associated with molecular hydrophobicity and is closely linked to biological activity.42 The partial correlation coefficients between all studied descriptors were below 0.70 (aij < 0.70), indicating that the selected descriptors are independent and do not exhibit significant multicollinearity.43
Predictive power estimation
The reliability of a QSAR model was evaluated using R², standard deviation (S), Fischer coefficient (F), and cross-validation coefficient (Q²CV). R², S, and F assess agreement between predicted and experimental values, 44,45 while Q2CV evaluates robustness and external predictivity. A model is considered reliable when data points closely align with the regression line.46 The fit of the points to the line can be evaluated by the coefficient of determination.
where
Yi,exp: Experimental antifungal activity value
yi,theo: Predicted antifungal activity value
yi,exp: Mean experimental antifungal activity value
Values of R² approaching 1 indicate strong correlation between experimental and predicted activities. The variance σ2 is calculated as:
where k is the number of descriptors, n is the number of molecules, and n-k-1 represents the degrees of freedom. The standard deviation (S), derived from σ², quantifies the precision of the model:
The statistical significance of the model is further evaluated by the Fischer coefficient (F):
Finally, the cross-validation coefficient (Q²CV) estimates the predictive accuracy on the validation set:
According to Eriksson et al.,47 a satisfactory model requires Q2CV ˃ 0.5, while values above ˃ 0.9 indicate excellent predictive power. In addition, the criterion R2-Q2CV ˂ 0.3 must be met to confirm model robustness. External predictive ability can also be assessed by comparing the ratio of predicted to experimental activities (log(1/C) theo/log(1/C) exp). The model is considered acceptable when this ratio approaches unity.
Acceptance Criteria for a QSAR Model
The performance of a QSAR model can be assessed by several complementary validation criteria. According to Eriksson et al.,47 a satisfactory model is characterized by a cross-validation correlation coefficient Q2CV > 0.5, while values of Q2CV > 0.9 indicate an excellent model. In addition, for a model to be considered robust, the difference between the determination coefficient and the cross-validation coefficient must satisfy the condition R2– Q2CV < 0.3.
For external validation, Tropsha et al,48,49 have proposed five widely accepted statistical criteria to evaluate predictive power. These criteria are:
Here, RTest2, R0‘2 are coefficients of determination for regressions through the origin with experimental versus predicted, and predicted versus experimental values, respectively.
Further refinement of QSAR validation criteria has been proposed by Roy and Roy,50 who introduced two metrics rm2 and ∆rm2, that measure the closeness between experimental and predicted values:
A QSAR model is considered acceptable only if both criteria are satisfied. These complementary approaches ensure a rigorous evaluation of both internal consistency and external predictive performance.
Applicability Domain (AD)
The final step in model development is to define the domain of applicability (AD), which establishes the chemical and biological space where predictions can be considered reliable.42, 51 This prevents risky extrapolations beyond the model’s training domain and ensures that predictions are limited to compounds similar in structure and activity to those used in model development.51,52 The AD was evaluated using Cook’s distance (Dᵢ), which quantifies the influence of each data point (even suspect points (outliers) in the results) on the regression,53,54 and given by
Where ȳ and ȳi are the 𝑛×1 vectors of predicted observations with and without the i-th data point, respectively, k is the number of model parameters, and is the σ2 variance of the regression. Di > 4/(n – k – 1), where n is the number of experimental points. The limit is determined by Cook’s distance, which is expressed as 4/((n-p-1). A compound is considered an outlier if its Cook’s distance exceeds the threshold.
Results and Discussion
Structural Characterization
Compounds 7a-j were synthesized via a three-step sequence, starting with the preparation of the key intermediate 1,3-thiazolidin-4-one (5). This intermediate 5 was obtained in two steps through the cyclocondensation of hydrazide (3) with mercaptoacetic acid 4 (Figure 2).55 In the next step, 1,3-thiazolidin-4-one (5) was condensed with a series of aromatic aldehydes (6a-j), affording the desired 1,3-thiazolidin-4-one-5-arylidene derivatives (7a–j). The products were isolated in good yields and purified by recrystallization from ethanol (Figure 3).
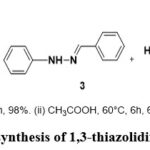 |
Figure 2: synthesis of 1,3-thiazolidin-4-one 5. |
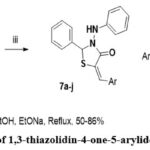 |
Figure 3: synthesis of 1,3-thiazolidin-4-one-5-arylidene derivatives (7a-j). |
Compounds 7a–j were synthesized and evaluated through structural characterization, biological testing, and QSAR analysis. Structural confirmation was achieved using 1H and 13C NMR spectroscopy, along with HRMS analysis. 1H NMR analysis confirmed successful condensation by the disappearance of the methylene (CH2) signals and the appearance of a new singlet around 8.0 ppm, corresponding to the vinylic proton (CH=) of the benzylidene moiety, consistent with the expected product. The aromatic region also displayed increased complexity, consistent with the introduction of additional aromatic substituents. The 13C NMR spectra further these findings, showing the disappearance of the methylene carbon resonance (around 29 ppm) and the emergence of a new signal near 137 ppm, attributable to the sp²-hybridized carbon of the CH=C group. Together with HRMS data, these results confirmed the successful synthesis of the target derivatives.
Antifungal and structure-activity relationships
Compounds 7a–j were tested against a clinical Aspergillus flavus using Itraconazole as the reference drug. Minimum Inhibitory Concentrations (MICs), initially expressed in µg/mL, were converted to micromolar (µM) to normalize differences in molecular weight among the compounds. This approach provides a more reliable comparison of antifungal efficacy by reflecting the number of active molecules present in solution. The MIC values of the synthesized derivatives are reported in Table 1.
Table 1: Antifungal activity (MIC, µM) of compounds 7a-j
| Compounds | Molecular Weight (g/mol) | MIC (µg/mL) | MIC (µM) |
| 7a | 358.4590 | 12.5 | 34.871 |
| 7b | 401.5280 | 6.25 | 15.565 |
| 7c | 374.4580 | 25 | 66.761 |
| 7d | 403.4560 | 25 | 61.964 |
| 7e | 376.4494 | 25 | 66.410 |
| 7f | 388.4850 | 50 | 128.705 |
| 7g | 427.3430 | >100 | >234.004 |
| 7h | 418.5110 | 100 | 238.942 |
| 7i | 437.3550 | >100 | >228.647 |
| 7j | 364.4810 | 25 | 68.590 |
| Itraconazole | 705.64 | 0.781 | 1.10 |
The results obtained showed that compounds (7a-j) exhibited moderate to strong antifungal activity against the clinical strain of Aspergillus flavus, with MICs ranging from 15.565 µM to 238.942 µM. The unsubstituted phenyl derivative 7a displayed moderate activity (34.871 µM), which may be attributed to the thiazolidinone ring and the phenylpropenone chain, a motif also found in chalcones known for antifungal properties.56,57
Compound 7b, bearing a para-dimethylamine substituent, emerged as the most active derivative with an MIC of 15.565 µM, approximately twice as potent as the unsubstituted phenyl analogue 7a (34.871 µM). This suggests that strong electron-donating (by mesomerism) groups at the para position enhance affinity for the biological target, likely through increased electronic interactions. By contrast, replacing the dimethylamine with a hydroxyl group (7c, MIC = 66.761 µM) or a methoxy group (7f, MIC = 128.705 µM) resulted in a sharp decline in activity, confirming that weaker electron-donating substituents are less favorable. Introducing two methoxy groups (7h, MIC = 238.942 µM) completely abolished activity. These results indicate that modulation of antifungal potency by electron-donating groups is highly sensitive to substituent strength and position, with only the dimethylamine substituent in the para position being beneficial.
Electron-withdrawing groups were also explored. The para-nitro-substituted derivative 7d displayed moderate activity (61.964 µM), but significantly weaker than 7b. Halogen substitution also failed to enhance activity: the para-chloro (7e, 66.410 µM), di-chloro (7g, 238.942 µM), and meta-bromo (7i, >228.647 µM) derivatives all showed reduced or negligible activity. This observation contrasts with antifungal azoles, where halogenation often enhances performance, suggesting a different structure–activity relationship in thiazolidinones. Due to the high structural diversity of chlorinated compounds and the limited exploration of less active congeners, largely for economic reasons, consistent rules describing the impact of chlorine substituents on biological properties remain elusive. This limitation, emphasized in the literature,58-60 underlines the scarcity of systematic data in the public domain. Within the scope of thiazolidinone derivatives, such complexity suggests that chlorine substitution may exert context-dependent effects on antifungal activity, warranting careful evaluation in SAR studies.
Finally, replacing the phenyl ring of 7a with a thiophene ring (7j, 68.590 µM) did not improve efficacy, yielding activity comparable to hydroxyl- and fluoro-substituted analogues. SAR analysis revealed that antifungal activity is primarily governed by the electronic nature of the para substituent, with the dimethylamine group showing the greatest potency.
QSAR Modeling
To rationalize these results, a QSAR study was conducted using molecular descriptors derived from quantum chemical calculations (Table 2).
Table 2: Molecular descriptors and antifungal activities
| Compounds | μD (Debye) | Pol (u.a) | EHOMO (ev) | ELUMO (eV) | ΔE (eV) | μ (eV) | η (eV) | LogP | EI (eV) | ω (eV) | χ (ev) | AE (ev) | |
| 7a | 4.457 | 1.749 | 323.724 | -0.217 | -0.078 | 0.139 | -0.148 | 0.069 | 4.820 | 0.217 | 0.157 | 0.148 | 0.078 |
| 7b | 4.807 | 2.648 | 386.402 | -0.186 | -0.058 | 0.128 | -0.122 | 0.064 | 4.800 | 0.186 | 0.117 | 0.122 | 0.058 |
| 7c | 4.176 | 3.838 | 335.310 | -0.206 | -0.067 | 0.139 | -0.137 | 0.069 | 4.590 | 0.206 | 0.134 | 0.137 | 0.067 |
| 7d | 4.207 | 10.310 | 360.753 | -0.231 | -0.111 | 0.120 | -0.171 | 0.060 | 3.250 | 0.231 | 0.244 | 0.171 | 0.111 |
| 7e | 4.177 | 4.041 | 321.340 | -0.217 | -0.074 | 0.143 | -0.146 | 0.072 | 4.830 | 0.217 | 0.149 | 0.146 | 0.074 |
| 7f | 3.890 | 2.501 | 349.920 | -0.210 | -0.067 | 0.142 | -0.138 | 0.071 | 4.930 | 0.210 | 0.134 | 0.138 | 0.067 |
| 7g | 3.622 | 4.069 | 310.370 | -0.225 | -0.086 | 0.139 | -0.155 | 0.070 | 6.030 | 0.225 | 0.174 | 0.155 | 0.086 |
| 7h | 3.604 | 4.232 | 363.980 | -0.204 | -0.061 | 0.143 | -0.132 | 0.071 | 4.880 | 0.204 | 0.122 | 0.132 | 0.061 |
| 7i | 3.637 | 1.402 | 342.520 | -0.224 | -0.082 | 0.142 | -0.153 | 0.071 | 5.600 | 0.224 | 0.164 | 0.153 | 0.082 |
| 7j | 4.164 | 3.467 | 318.080 | -0.010 | -0.213 | -0.203 | -0.112 | -0.102 | 4.380 | 0.010 | -0.062 | 0.112 | 0.213 |
Three descriptors: dipole moment (µD), polarizability (Pol), and lipophilicity (LogP), emerged as key contributors to antifungal activity. The best regression model (Equation 2) demonstrated strong predictive power (R² = 0.993; Q²CV = 0.952), indicating that these descriptors reliably capture the structural determinants of activity:
pCL100 = 12.36677 – 0.11148 * µD (Debye) – 1.25810 * 10-2 * Pol(u.a) – 0.76371 * LogP
N = 7; R2 = 0.993; Q2CV = 0.952; S = 0,622; F = 134.003; R2 – Q2CV = 0.041
According to this model, low dipole moment, low polarizability, and reduced lipophilicity favor stronger antifungal activity. In particular, polarity emerged as a critical factor, with highly polar molecules showing weaker inhibition of A. flavus.
The values of the partial correlation coefficients of the descriptors (aij) in the model, external validation pCL100Theo / pCL100Exp are shown in Table 3 and Table 4, respectively.
Table 3: Correlation matrix (Pearson(n)) between the different descriptors
| Variables | pCL100Exp | µD (Debye) | Pol (u.a) | LogP |
| pCL100Exp | 1 | 0.022 | 0.298 | -0.475 |
| µD (Debye) | 0.022 | 1 | 0.186 | -0.716 |
| Pol (u.a) | 0.298 | 0.186 | 1 | -0.342 |
| LogP | -0.475 | -0.716 | -0.342 | 1 |
Values in bold are significantly different from 0 at p < 0.05. Highly significant at p < 0.01.
Table 4: Experimental and theoretical activities and their relationship with compounds in the validation set
| Compounds | pCL100Exp | pCL100Theo | pCL100Theo / pCL100Exp |
| 7b | 4.807 | 3.544 | 0.7 |
| 7g | 3.622 | 3.403 | 0.9 |
| 7j | 4.164 | 4.633 | 1.1 |
External validation using compounds 7b, 7g and 7j confirmed the robustness of the model (Table 4). The ratio pCL100Theo / pCL100Exp of theoretical to experimental activity values was close to unity (1) for all three compounds, indicating a good agreement between predicted and observed results. The regression line of experimental versus theoretical activity (Figure 4) further supported the reliability of the QSAR model.
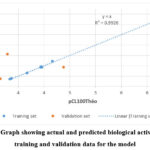 |
Figure 4: Graph showing actual and predicted biological activity for all training and validation data for the model |
Model Validation
The robustness and predictive reliability of the developed QSAR model were assessed using Tropsha’s and Roy’s criteria.
Verification of Tropsha’s Criteria
All conditions were satisfied, confirming that the model is statistically acceptable for predicting antifungal activity. Verification according to Roy’s criteria further supported this conclusion
Verification Roy’s Criteria
With rm2 = 0.950 and rm‘2 = 0.806
Thus, both Tropsha’s and Roy’s validations confirm the acceptability of the QSAR model.
The model identified three key descriptors as determinants of antifungal activity: dipole moment (µD), lipophilicity (LogP), and polarizability (Pol). Among these, lipophilicity contributed most strongly to activity prediction, followed by dipole moment and polarizability (Figure 5). Lower polarity and reduced lipophilicity were found to favor improved activity.
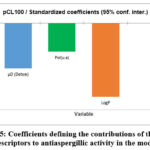 |
Figure 5: Coefficients defining the contributions of the three descriptors to antiaspergillic activity in the model. |
Among the model descriptors, lipophilicity is the most important.
The predictive performance of the model was further assessed through similarity curves comparing experimental and predicted values (Figure 6).
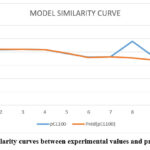 |
Figure 6: Similarity curves between experimental values and predicted values. |
Applicability domain
Apart from a single deviation point, the data showed minimal error, confirming the reliability of the model for activity prediction.
The applicability domain was determined using Cook’s distance (Figure 7).
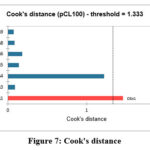 |
Figure 7: Cook’s distance. |
Results indicated that compound 7a is an outlier and should be excluded from predictive applications. However, for all other compounds (7b–j), the descriptors exerted a strong structural influence on the model, validating their suitability for reliable activity prediction
Overall, the QSAR analysis revealed that antifungal activity in this series is primarily governed by lipophilicity, dipole moment, and polarizability. Low polarity and controlled lipophilicity were essential for optimal performance. With the exception of compound 7a, all derivatives showed strong influence on the model. These findings provide valuable guidance for designing new thiazolidinone derivatives with improved antifungal activity, paving the way for the development of more effective antifungal agents.
Conclusion
This study reports the successful design, synthesis, and antifungal evaluation of novel 1,3-thiazolidin-4-one-5-arylidene derivatives. Structural characterization by 1H/13C NMR and HRMS confirmed the synthesized compounds. Biological testing against a clinical Aspergillus flavus strain revealed activity dependent on the electronic nature and substitution pattern of the aryl ring at position 5, with compound 7b (para-dimethylamine) showing the highest potency (MIC = 15.56 µM). QSAR analysis identified lipophilicity, dipole moment, and polarizability as key descriptors, with lipophilicity being the primary factor enhancing antifungal activity. The validated QSAR model demonstrated strong predictive performance, providing a reliable framework for guiding future structural modifications. These findings underscore the thiazolidinone scaffold as a promising platform for developing next-generation antifungal agents and offer valuable design principles for optimizing biological efficacy.
Acknowledgement
The authors thank Angré University Hospital Center (Côte d’Ivoire) for antifungal testing and Université de Nantes, CiMed-AE1155, France, for providing materials and facilities for spectroscopic analyses.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
References
- Gangneux, J.P.; Bouchara, J.P.; Chabasse, D. Biologie et diagnostic des infections à Aspergillus EMC – Pédiatrie – Maladies infectieuses. 2013, 30(4), 1-10.
- Andrès, E.; Tiphine, M.; Letscher-Bru, V.; Herbrecht, R. Nouvelles formes lipidiques de l’amphotéricine B. Revue de la littérature. La Revue de Médecine Interne. 2001, 22(2),141-150.
CrossRef - Domin, J.P. Le Programme de médicalisation des systèmes d’information (PMSI). Histoire, médecine et santé. 2013, (4), 69-87.
CrossRef - Guillaume, D.; Chandenier, J. Aspergillus_and_aspergillus_diseases. Feuillets de Biologie. 2025, 51(293),53-63
- Latgé, J.P. ; Chamilos G. Aspergillus fumigatus and Aspergillosis in 2019. Clinical Microbiology Reviews 2019, 33(1), e00140-18.
CrossRef - Bottery, M. J.; van Rhijn, N.; Chown, H.; Rhodes, J. L.; Celia-Sanchez, B. N.; Brewer, M. T.; Momany, M.; Fisher, M. C.; Knight, C. G.; Bromley, M. J. Elevated mutation rates in multi-azole resistant Aspergillus fumigatus drive rapid evolution of antifungal resistance. Nature Communications 2024, 15(1), 1–12.
CrossRef - Latgé, J.P. Aspergillus fumigatus and Aspergillosis. Clinical Microbiology Reviews 1999, 12(2), 310-350.
CrossRef - Vandeputte, P; Ferrari, S.; Coste, A.T. Antifungal Resistance and New Strategies to Control Fungal Infections. International Journal of Microbiology. 2012, 2012, 1-26.
CrossRef - Perlin, D.S. Echinocandin Resistance in Candida. Clinical Infectious Diseases. 2015, 61(suppl_6), S612-S617.
CrossRef - Williams, C.; Ramage, G. Fungal Biofilms in Human Disease. In: Donelli G, ed. Biofilm-Based Healthcare-Associated Infections. Vol 831. Advances in Experimental Medicine and Biology. Springer International Publishing. 2015, 11-27.
CrossRef - Borghi, E; Morace, G.; Borgo, F; Rajendran, R; Sherry, L.; Nile, C.; Gordon, R. New strategic insights into managing fungal biofilms. Frontiers in Microbiology 2015, 6, 1077.
CrossRef - Sharma, R.; Kumar, K.; Singh, K.; Joshi, R. Shared Based Rate Limiting: An ISP level Solution to Deal DDoS Attacks. In: 2006 Annual IEEE India Conference. IEEE. 2006, 1-6.
CrossRef - Ottanà, R.; Maccari, R.; Barreca, M.L.; Bruno, G.; Rotondo, A.; Rossi, A., Giuseppa, C.; Rosanna, D. P.; Lidia, S.; Salvatore, C.; Maria, G. V. 5-Arylidene-2-imino-4-thiazolidinones: Design and synthesis of novel anti-inflammatory agents. Bioorganic & Medicinal Chemistry. 2005, 13(13), 4243-4252.
CrossRef - Goel, B.; Ram, T; Tyagi, R.; Bansal, E.; Kumar, A.; Mukherjee, D.; Sinha, J. 2-Substituted-3-(4-bromo-2-carboxyphenyl)-5-methyl-4-thiazolidinones as potential anti-inflammatory agents. European Journal of Medicinal Chemistry. 1999; 34(3), 265-269.
CrossRef - Vicini, P.; Geronikaki, A.; Anastasia, K; Incerti, M.; Zani, F. Synthesis and antimicrobial activity of novel 2-thiazolylimino-5-arylidene-4-thiazolidinones. Bioorganic & Medicinal Chemistry. 2006, 14(11), 3859-3864.
CrossRef - Bondock, S.; Khalifa, W.; Fadda, A.A. Synthesis and antimicrobial evaluation of some new thiazole, thiazolidinone and thiazoline derivatives starting from 1-chloro-3,4-dihydronaphthalene-2-carboxaldehyde. European Journal of Medicinal Chemistry. 2007, 42(7), 948-954.
CrossRef - Küçükgüzel, G.; Kocatepe, A; De Clercq, E.; Şahin, F.; Güllüce, M. Synthesis and biological activity of 4-thiazolidinones, thiosemicarbazides derived from diflunisal hydrazide. European Journal of Medicinal Chemistry. 2006, 41(3), 353-359.
CrossRef - Fahmy, H.T. Synthesis of some new triazoles as potential antifungal agents. Bollettino chimico farmaceutico 2001, 140(6), 422-427.
- Solomon, V.R.; Haq, W. Srivastava, K.; Puri, S.K.; Katti, S.B. Synthesis and Antimalarial Activity of Side Chain Modified 4-Aminoquinoline Derivatives. Journal of Medicinal Chemistry 2007, 50(2), 394-398.
CrossRef - Shah, T.J.; Desai, V.A. Synthesis of some novel fluorinated 4-thiazolidinones containing amide linkages and their antimicrobial screening. Arkivoc. 2007, 2007(14), 218-228.
CrossRef - Kavitha, C.V.; Basappa; Swamy, S.N; Mantelingu, K.; Doreswamy, S., Sridhar, M.A.; J. Shashidhara, P.; Kanchugarakoppal, S. R. Synthesis of new bioactive venlafaxine analogs: Novel thiazolidin-4-ones as antimicrobials. Bioorganic & Medicinal Chemistry. 2006, 14(7), 2290-2299.
CrossRef - Babaoglu, K.; Page, M.A.; Jones, V.C; McNeil, M.R.; Dong, C.; Naismith, J.H.; Lee, R.E. Novel inhibitors of an emerging target in Mycobacterium tuberculosis; substituted thiazolidinones as inhibitors of dTDP-rhamnose synthesis. Bioorganic & Medicinal Chemistry Letters. 2003, 13(19),3227-3230.
CrossRef - Rahman, V.P.M; Mukhtar, S.; Ansari, W.H.; Lemiere, G. Synthesis, stereochemistry and biological activity of some novel long alkyl chain substituted thiazolidin-4-ones and thiazan-4-one from 10-undecenoic acid hydrazide. European Journal of Medicinal Chemistry. 2005, 40(2):173-184.
CrossRef - Yamanouchi, T. Concomitant therapy with pioglitazone and insulin for the treatment of type 2 diabetes. Vascular Health and Risk Management 2010, 6, 189-197.
CrossRef - He, L.; Liu, X.; Wang, L.; Yang, Z. Thiazolidinediones for nonalcoholic steatohepatitis: A meta-analysis of randomized clinical trials. Medicine (Baltimore). 2016, 95 (42), e4947.
CrossRef - Parisa, B.; Alborzi, A.; Pedram, H.; Mahsa, M. Antifungal susceptibility of the Aspergillus species by Etest and CLSI Reference Methods., Archives of Iranian Medicine. 2012, 15(7), 429-32
- Chatterjee, S.; Ali, S. H. Simple linear regression. Regression Analysis by Example, John Wiley & Sons. Inc., Hoboken, 2006.
CrossRef - Huynh, T.N.P. Synthèse et études des relations structure/activité quantitatives (QSAR/2D) d’analyse benzo[c]phénanthridiniques. PhD thesis. Université d’Angers; 2007. https://theses.hal.science/tel-00346332.
- Espinel-Ingroff, A.; Arthington-Skaggs, B.; Iqbal, N.; Ellis, D.; Pfaller, M.A.; Messer, S.; Rinaldi, M.; Fothergill, A.; Gibbs, D. L.; Wang, A.; Multicenter Evaluation of a New Disk Agar Diffusion Method for Susceptibility Testing of Filamentous Fungi with Voriconazole, Posaconazole, Itraconazole, Amphotericin B, and Caspofungin. Journal of Clinical Microbiology. 2007, 45(6), 1811-1820.
CrossRef - Cantón, E.; Espinel-Ingroff, A.; Pemán, J. Trends in antifungal susceptibility testing using CLSI reference and commercial methods. Expert Review of Anti-infective Therapy. 2009, 7(1), 107-119.
CrossRef - Arikan, S. Current status of antifungal susceptibility testing methods. Medical Mycology. 2007, 45(7), 569-587. doi:10.1080/13693780701436794
CrossRef - Lass‐Flörl, C.; Perkhofer, S. In vitro susceptibility‐testing in Aspergillus species. Mycoses. 2008, 51(5), 437-446.
CrossRef - Gaussian 09, Revision A.02, Frisch, M. J.; Trucks, G. W.; Schlegel, H. B.; Scuseria, G. E.; Robb, M. A.; Cheeseman, J. R.; Scalmani, G. ; Barone, V. ; Mennucci, B.; Petersson, G. A.; Nakatsuji, H.; Caricato, M.; Li, X.; Hratchian, H. P.; Izmaylov, A. F.; Bloino, J.; Zheng, G. ; Sonnenberg, J. L.; Hada, M.; Ehara, M.; Toyota, K.; Fukuda, R.; Hasegawa, J.; Ishida, M.; Nakajima, T.; Honda, Y.; Kitao, O.; Nakai, H.; Vreven, T.; Montgomery, Jr.; Peralta, J. E.; Ogliaro, F.; Bearpark, M.; Heyd, J. J.; Brothers, E.; Kudin, K. N.; Staroverov, V. N.; Kobayashi, R.; Normand, J.; Raghavachari, K.; Rendell, A.; Burant, J. C.; Iyengar, S. S.; Tomasi, J.; Cossi, M.; Rega, N.; Millam, J. M.; Klene, M.; Knox, J. E.; Cross, J. B.; Bakken, V. ; Adamo, C.; Jaramillo, J.; Gomperts, R.; Stratmann, R. E.; Yazyev, O.; Austin, A. J.; Cammi, R.; Pomelli, C.; Ochterski, J. W.; Martin, R. L.; Morokuma, K.; Zakrzewski, V. G.; Voth, G. A.; Salvador, P.; Dannenberg, J. J.; Dapprich, S. ; Daniels, A. D.; Farkas, O.; Foresman, J. B.; Ortiz, J. V.; Cioslowski, J.; Fox, D. J. Gaussian, Inc., Wallingford CT, 2009.
- Chattaraj, P.K.; Cedillo, A.; Parr, R.G. Variational method for determining the Fukui function and chemical hardness of an electronic system. The Journal of Chemical Physics. 1995, 103(17), 7645-7646.
CrossRef - Ayers, P.W.; Parr, R.G. Variational Principles for Describing Chemical Reactions. Reactivity Indices Based on the External Potential. Journal of the American Chemical Society. 2001, 123(9), 2007-2017.
CrossRef - De Proft, F; Martin, J.M.L.; Geerlings, P. On the performance of density functional methods for describing atomic populations, dipole moments and infrared intensities. Chemical Physics Letters. 1996, 250(3-4), 393-401.
CrossRef - De Proft, F.; Martin, J.M.L.; Geerlings, P. Calculation of molecular electrostatic potentials and Fukui functions using density functional methods. Chemical Physics Letters. 1996, 256(4-5), 400-408.
CrossRef - Microsoft Excel 2016 : Guide étape par étape https://fr.wps.com/articles-excel/fr-microsoft-excel-2016-free-download/.
- XLSTAT version 2021.2.2. XLSTAT, Your data analysis solution. Published June 4, 2021. https://www.xlstat.com/news/xlstat-version-2021-2-2.
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Advanced Drug Delivery Reviews 2001, 46(1-3), 3-26.
CrossRef - Xavier, S; Periandy, S.; Ramalingam, S. NBO, conformational, NLO, HOMO–LUMO, NMR and electronic spectral study on 1-phenyl-1-propanol by quantum computational methods. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy. 2015, 137, 306-320.
CrossRef - Chtita, S. Modélisation de molécules organiques hétérocycliques biologiquement actives par des méthodes QSAR/QSPR. Recherche de nouveaux médicaments. phdthesis. Université Moulay Ismaïl, Meknès ; 2017. https://hal.science/tel-01568788.
- Vessereau A. Methodes statistiques en biologie et en agronomie, 1988, ISBN 2-85206-834-6.
- Snedecor, G.W. and Cochran, W.G. Statistical Methods. Oxford and IBH, New Delhi, 1967, 381-418.
- Diudea, M.V. QSPR/QSAR Studies for Molecular Descriptors. Nova Science, Huntingdon, New York. 2000
- Esposito, E.X.; Hopfinger, A.J.; Madura, J.D.. Methods for Applying the Quantitative Structure-Activity Relationship Paradigm. In: Bajorath, J. (eds) Chemoinformatics. Methods in Molecular Biology 2004, 275, 131-213.
CrossRef - Eriksson, L.; Jaworska, J.; Worth, A.P.; Cronin, M.T.D.; McDowell, R.M.; Gramatica, P. Methods for reliability and uncertainty assessment and for applicability evaluations of classification- and regression-based QSARs. Environmental Health Perspectives. 2003, 111(10), 1361-1375.
CrossRef - Golbraikh, A.; Tropsha, A. Beware of q2! Journal of Molecular Graphics and Modelling. 2002, 20(4), 269-276.
CrossRef - Tropsha, A.; Gramatica, P.; Gombar, V.K. The Importance of Being Earnest: Validation is the Absolute Essential for Successful Application and Interpretation of QSPR Models. QSAR & Combinatorial Science 2003, 22(1), 69-77.
CrossRef - Roy, P.P.; Roy, K. On Some Aspects of Variable Selection for Partial Least Squares Regression Models. QSAR & Combinatorial Science 2008, 27(3), 302-313.
CrossRef - Ghamali, M.; Chtita, S.; Bouachrine, M.; Lakhlifi, T. Méthodologie générale d’une étude RQSA/RQSP. Revue Interdisciplinaire. 2016, 1(1). https://revues.imist.ma/index.php/Revue-Interdisciplinaire/article/view/6222.
- Prana, V. Approches structure-propriété pour la prédiction des propriétés physico-chimiques des substances chimiques. UPMC-Ineris, Paris. Published online 2013. http://vinca.prana.free.fr/ These_VP.pdf.
- Dı́az-Garcı́a, J.A.; González-Farı́as, G. A note on the Cook’s distance. Journal of Statistical Planning and Inference. 2004, 120(1-2), 119-136.
CrossRef - Militino, A. F.; Palacios, M.B.; Ugarte, M.D. Outliers detection in multivariate spatial linear models. Journal of Statistical Planning and Inference. 2006, 136(1), 125-146.
CrossRef - Camara, T.E; Koné, A.; Coulibaly, B.; Kra, A.S.; Coulibaly, P. M. A.; Coulibaly, S.; Ballo, D.; Coulibali, S. Solvent-free synthesis of novel 1,3-thiazolidin-4-one-5-arylidene derivatives via cyclocondensation. Journal of Sulfur Chemistry. 2025, 1-11.
CrossRef - Omar, K.; Geronikaki, A.; Zoumpoulakis, P.; Camoutsis, C.; Soković, M.; Ćirić, A.; Jasmina, G. Novel 4-thiazolidinone derivatives as potential antifungal and antibacterial drugs. Bioorganic & Medicinal Chemistry. 2010, 18(1), 426-432.
CrossRef - Adhikari, S.; Nath, P.; Deb, V.K.; Das, N.; Banerjee, A.; Pathak, S.; Asim, K. D.; Pharmacological potential of natural chalcones: a recent studies and future perspective. Front Pharmacol. 2025, 16, 1570385.
CrossRef - Dietrich, H., Angew. Chem. 1994, 106, 1997.
- Dietrich, H. Toxicity of Chlorinated Organic Compounds: Effects of the Introduction of Chlorine in Organic Molecules. Angew. Chem. Int. Ed. Engl., 1994, 33, 1920-1935.
CrossRef - Šegan, S.; Krunić, M. J.; Andrić, D. B.; Šukalović, V. B.; Penjišević, J. Z.; Jevtić, I. I. Evaluation of lipophilicity and drug-likeness of donepezil-like compounds using reversed-phase thin-layer chromatography. Biomedical Chromatography, 2024, 38(7), e5867.
CrossRef
Accepted on: 08 Oct 2025
Second Review by: Dr. Rafid Saad Dawood
Final Approval by: Dr. Abdelwahab Omri