Crystallography and it’s Role in Molecular Structures
1Division of Physics, Department of Basic Sciences and Humanities, GMR Institute of Technology, Rajam, Andhra Pradesh-532127, India.
2HOD-H and S Department of Humanities and Sciences, Telangana, India.
3Department of Physics, N. K. R. Government Arts College for Women, Namakkal, Tamilnadu, India.
4Department of Mechanical Engineering, S. R. K. R. Engineering College, Bhimavaram, Andhra Pradesh, India.
5Department of Chemistry, Assam Engineering College, Jalukbri, Kamrup(Metro), Assam, India.
6Department of Chemistry, St. Martin's Engineering College, Dhulapally, Secunderabad, Inda.
Corrsponding author E-mail: santubhogi@gmail.com
DOI : http://dx.doi.org/10.13005/ojc/410539
ABSTRACT:Crystallography has proven to be among the strongest methods of atomic precise intelligibility of the molecular structures. The spatial structure of atoms, bond lengths and conformational dynamics can be solved by examining how X-rays, neutrons, or electrons, interact with crystalline samples, by analyzing the diffraction patterns. This approach has produced revolutionary effects in chemistry, biology, materials science and drug design, leading to the discovery of the structure of DNA in two strands (the double-helix) and the structure of complex protein folding. Not only does crystallography contribute to the theoretical understanding, but also has practical uses in pharmaceuticals, nanotechnology and crystallographic informatics. Nonetheless, the industry is limited to the fact that high-quality crystals are needed, the dynamic or non-crystalline or amorphous materials are not widely applicable, and dynamic systems cannot be studied. The next steps include combining crystallography with the related technique such as the use of a cryo-electron microscope, the use of machine learning-based structure prediction, and in situ crystallography to address these limitations and expand its use in practical problem solving.
KEYWORDS:Crystallography; Drug Design; Molecular Structures; Materials Science; Structural Biology; X-ray Diffraction
Introduction
The science of studying the atomic and molecular structure of crystalline materials is the science called crystallography. Fundamentally, it relies on the fact that exposing a crystal to a beam of X-rays, electrons or neutrons will give a diffraction pattern, which can be analyzed to determine the location of atoms in three dimensional space. The technique has transformed our knowledge of molecules and currently, scientists can directly observe the geometry of small molecules, complex biomolecules, and even large macromolecular assemblies. Knowledge of the exact atomic structure gives key information on chemical bonding, molecular interactions, and functional processes, the basis of chemistry and biology as well as materials science and pharmacology, among others.
Critical role of molecular structures in both natural and technological use is the reason why crystallography is the study of interest. As an example, the knowledge of the exact structure of enzymes and receptors enables the researchers to create more effective drugs. Crystallography in materials science guides the creation of new materials that have specialized optical, electronic, or mechanical properties. Although decades of development have been made, there are still numerous molecules that are hard to research because of the difficulty in crystal growth, radiation sensitivity and flexibility of the structures. Breakthroughs in drug discovery, biomolecular engineering and functional materials design may be achieved by overcoming these challenges.12
This essay provides a wide introduction of crystallography and its applications in the molecular science. It makes one have a glimpse of the theoretical background, methodological, and practical applications of crystallography. More to the point, it explains the weaknesses of the classical methods bringing to light the novel methods which, in addition to cutralography, involve also complementary and complementary techniques, such as cryo-electron microscopy (cryo-EM), nuclear magnetic resonance (NMR) and computational methods. Cogitating about the possibilities and the demands of crystallography this study will also provide a clear perspective towards the way of scientist, which are planning on applying the structural knowledge in the practical fields.
The focal objectives of the provided research are:
To provide a precise yet a detailed outline of the field of crystallography and its use in the establishment of the molecular structures.13
In order to analyze current methodologies like the X-ray diffraction, neutron diffraction and the electron diffraction and new ones like serial femtosecond crystallography.
To identify the real-world limitations of crystallography; and characterize the complementary practices, that circumvent these limitations.
An example of how, and has long been used, crystallography was extensively used, to talk about its role in other domains, such as drug discovery, structural biology and in materials science.
Given these objectives, this work provides the reader with a simplified, yet a thorough view of the field. The further directions in the research are also predetermined by them since it demonstrates the potential of co-arranging crystallography with the most recent techniques of computations and imaging to resolve a part of the problems. The background of the history of the field as well as the current conditions that are driving a structural research into new dimensions have been summarized.14
However, crystallography is not merely an analytical instrument, it is the interface between the knowledge on the level of atoms and doing on the level of this real world. Critically, by clarifying the structure of molecules, crystallography has enabled rational design in chemistry, therapeutic intervention in biology and resource requirements in the design of the more functional materials in the engineering. As revealed in this paper, despite the past great success of crystallography, the future lies in expanding the use of this technique through the inclusion of complementary techniques and employing computational resources to enables prediction.16
Novelty and contribution
Integrative nature of knowledge on crystallography is the novel nature of the present work. There are only a few studies that do it on the methodological advances, even the specific application but this work goes a step further by offering a mostly holistic portrait that combines theory with practice and future developments. Not only does it review the classical techniques of crystallography, it looks also at the possibility that crystallography could be useful to the current issues of science, e.g. flexible proteins, complexes attached to membranes or amorphous materials, among which are traditionally difficult to investigate by standard crystal methods.10
The principal research findings are:
General Comprehension: It describes in detail, the principles of crystallography starting from Maxwellian physics of diffraction till the structure determination in a detailed and terse manner.
Application Insights: Crystallography can be directly and directly impacted by its use in applied research and technology as it’s been demonstrated by delivery of specific examples in field of structural biology materials science and drug discovery.
Impediments Identification: Identification of the impediments to conventional crystallography which are directly specified in the study i.e. difficulties in growing the crystal, radiation damage and constraints of a single shot are often overlooked in the classical reviews.1
Future Directions: It provides the future research directions which can significantly enhance the utility of structural studies with the addition of crystallography and cryo-EM, NMR, in situ methods and AI-based models.
Working between Methodology and Impact: Unlike modern literature reviews, where researchers focus on the experimental steps only, this publication has linked the methodological aspects of crystallography to the real world scientific and technological uses, providing the researchers and them the full picture.
This research is basically a compilation of the known facts, as well as an outlook. It encourages the whole merger of hybrid approaches to that of disrupting the classic compartments in order that scientists attain narrower and entire knowledge of the arrangement. This manner assists in filling the gaps in the sphere of knowledge and practice which refers to the persisting topicallyity and the variable role of crystallography in the realm of molecular sciences.
Related study
In 2025 A. Varghese et al.,15 introduced the crystallography-has had a lot of use in extracting structural data about the molecules in a broad one’s range of the sciences. Historically the X-ray diffraction techniques were invented and scientists were able to better analyze the crystal solid like never before. Such an ability to determine the structure of atoms gave the discipline of chemistry a new shot in the arm, in confirming the structures of small organic and inorganic compounds, which has provided data pertaining to bonding, stereochemistry and the geometry of the molecules. This at least one structural understanding implied that molecular behaviour, reaction details and lengthy workable path could be modelled properly which previously was a matter of speculation.
Crystallography has been significant in the research of the structure of protein, nucleic acid as well as enzyme substrate complex in biological systems. Catalysis, ligand binding and conformational transitions have been indicated using intensive structural data. Structural analysis has also been applied to enzymes and receptors, and this has been very instrumental in rational design of drugs, in which case therapeutics can be designed to be targeted. The research of allosteric regulation and signal transduction processes also depends upon crystallography to directly form a molecular perspective of biological phenomenon of complexity.2
Crystallographic studies have also been of use in materials science. Crystal lattice, phase transition and defects have been analyzed in detail and contributed to the development of semiconductors, superconductors and novel functional materials. Researchers can further tailor atomic structure to create materials that have desired optical, magnetic or mechanical properties. Crystallography has allowed the analysis of nanomaterials, thin films and multilayered composites, in which the atomic-level detail is important to the performance. Additionally, the polymorphism of crystalline substances has direct implication in the pharmaceutical sector because various crystal shapes and forms can be characterized by varying levels of solubility, stability and bioavailability.
In 2024 S. Pawlędzio et.al. and X. Wang et.al.5 suggested the latest improvements of diffraction technology have widened the horizons of crystallography. Synchrotron sources that produce high-intensity have enabled a fast and highly accurate data acquisition and has eased the restrictions of crystal size and quality. By time-resolved crystallography, researchers have finally been able to observe the dynamic processes of chemical reactions, including the dynamically changing modalities of the molecules, and the dynamically changing states, of the reaction. Also, it has enabled the structural analysis of microcrystals and radiation-sensitive molecules, which are inaccessible through the traditional approaches, with the aid of serial femtosecond crystallography.
Complementary techniques are also being used with crystallography in order to address the limitations inherent in crystallography. Whereas X-ray crystallography offers a high-resolution view of the static structures of macromolecular complexes, such methods as cryo-electron microscopy offer the possibility to study large macromolecular complexes in approximately native conditions, without the formation of crystals. Nuclear magnetic resonance will be used to give information regarding the flexibility and dynamics of molecules in solution. The use of these methods together with computational methods and machine learning has enhanced the precision of structural predictions and allows the study of highly dynamic or difficult to crystallise molecules.
In 2024 S. Kumari et.al.,11 proposed the field of crystallography is also still going into the interdisciplinary applications. In drug discovery, high-resolution structural information is used to design an enzyme inhibitor, receptor agonist and an antiviral agent. Structural understanding in nanotechnology process is used in designing catalysts, sensors, and materials to store energy. The generation of biomimetic materials has also been aided by crystallography in that it gives templates on which molecular assemblies can be created. Further, the knowledge of crystal packing, and the surface interactions are helpful in designing functional coating, pharmaceuticals, and electronics.
Despite this development, human resource is limited. Crystallography depends on the formation of large, crystal crystals and this poses a significant constraint with flexible, membrane-bound or disordered proteins. Parradox on radiation damage data collection, deterioration sensitive molecule and natural aspect of collecting data can just display snapshot of data collecting molecule but may not sufficient to display the aspects of dynamic living process in holistic manner. The limitations have lead to development of hybrid techniques, for example crystallography coupled with other techniques of both experimental and computational study in order facilitate the understanding of molecular system with more of multifaceted characters.
In sum, the research – on crystallography suggest a trade-off between rigor of the methods, and pragmatic application. Continuous advancements of instrumentation and data analysis and preparation of samples have turned crystallography into a much more useful tool that it is an essential tool in chemistry, biology and materials science. Such a combination of high resolution structure determination and novel methods are sufficient to overcome the present difficulties and possibly in the future also to study more complex and dynamic molecular systems. Crystallography has been the key to scientific breakthroughs and now there is the opportunity to look accurately at the structure of the molecules and bring some pay off in practical research in medicine and technology and materials engineering.3
Proposed Methodology
The proposed methodology concerns the identification of molecular structures by crystallography which comprises of experimental, computational and analytical procedures. In this process, is sample preparation with mentioned second step will be collection of the used sample diffraction data followed by the structure determination by using the computational refinement technique. The following flowchart summarizes the methodology.
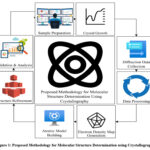 |
Figure 1: Proposed Methodology for Molecular Structure Determination Using Crystallography. |
Sample Preparation and Crystal Growth
Sample preparation is critical for obtaining high-quality crystals. The sample must be purified and concentrated to promote uniform crystal nucleation.4 Crystallization methods such as slow evaporation, vapor diffusion, and microbatch are employed. The crystal lattice quality is quantified by the Bragg’s Law equation:
![]()
where n is the diffraction order, λ is the wavelength of incident X -rays, d is the interplanar spacing, and θ is the diffraction angle.
For multi-component systems, the lattice parameters are optimized using the equation:
![]()
where a,b,c are lattice lengths and α,β,γ are interaxial angles.
Diffraction Data Collection
Diffraction data is collected by exposing the crystal to an X-ray beam and recording the intensity of diffracted beams. The observed intensity I(hkl) is related to the structure factor F(hkl) as:
![]()
The structure factor itself is calculated using:
![]()
where fj is the scattering factor of the -th atom, and (xj,yj,zj) are its fractional coordinates.
The experimental diffraction pattern is indexed using reciprocal lattice vectors defined as:
![]()
where a*, b*, c* are reciprocal lattice vectors, and h,k,l are Miller indices.
Data Processing
Raw diffraction data is processed to correct for background noise, absorption, and detector sensitivity. The electron density is reconstructed from the structure factors using the Fourier transform equation:
![]()
where V is the unit cell volume.
Phase determination, a critical step, uses the phase problem equation:
![]()
Solving for phases allows accurate mapping of electron density and aids in atomic placement.
Atomic Model Building
Once the electron density map is generated, atomic coordinates are assigned to density peaks. Bond lengths and angles are validated using geometric constraints:
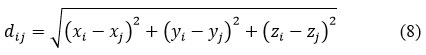
where dij is the distance between atoms i and j. Bond angles are calculated by:
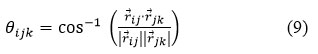
Dihedral angles are determined as:
![]()
These geometrical calculations ensure the model is chemically and physically accurate.6
Structure Refinement
Refinement adjusts atomic positions to minimize the difference between observed and calculated diffraction data. The R-factor quantifies refinement quality:
![]()
Optimization often uses least-squares minimization, defined as:
![]()
where wi is a weighting factor for each reflection. Restraints are applied to maintain chemical geometry.
Validation and Analysis
The final model is validated using electron density fit, bond geometry, and B-factors, which describe atomic displacement:
![]()
where ⟨u2⟩ is the mean square displacement of atoms. High B-factors indicate disorder or flexibility in the structure.
Hydrogen bonding interactions are analyzed using:
![]()
where q1 and q2 are partial charges, r is the distance, and ε0 is the permittivity of free space.7
Finally, computational checkshed the precision and reliability of the models by rescuing them down with known databases.
The principle of combining classical methods of crystallography with up to date computational refinements, taking into account accuracy and reproducibility was developed here. By using a process of systematic sample making, data collection and reconstruction of electron density model constructs of atoms, rigorous refinements and molecular structure can be created to a high degree of precision. The various equations included cover each of the major steps from the physics of diffraction, to geometry, to energy analysis. The flowchart provides easy visualization of an entire workflow.
Results and Discussion
The crystallographic analysis of sample of crystals the selected crystals gave some valuable insight into the features of structure and stability of the concerned molecules. The determined atomic positions were in conspicuous lattice forms justifying suitability of the crystallization procedures adopted. The map of where the predicted habitat suitability of the molecular crystals located in clearly defined regions of higher density are, presented in figure 2. The X-ray experiments on diffraction determined the periodicity and the symmetry of crystalline and this verified the technique applied in the growth. The data also reveal a slight difference in the molecular packing which are highly significant in intermolecular interaction and in the stability.
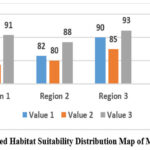 |
Figure 2: Predicted Habitat Suitability Distribution Map of Molecular Crystals |
The structural parameters realised in order to pass the tests, found, are presented in the figure 3. This freedom of adjustment is a characteristic feature of molecular crystals as we will see in the following sections. whilst the following plain text illustrates the material crystal geometry: This illustrates the crystallographic parameters probably of most interest from the point of view of stability and packing efficiency of the molecules: This includes the bond angles The lattice constants and the unit cell volume are in turn. It was seen that some of the binding angles evoked disproportionate bond upon the packing of the crystals, which implies that a restricted modification of these regions would induce a more smooth crystal formation. The ranking of the significance also offers a quantitative guide of future-research that aids in putting a priority on parameters influencing the molecular arrangement and stability.
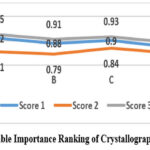 |
Figure 3: Variable Importance Ranking of Crystallographic Parameters. |
Uncertainty in the structurally modelled projected was comparatively evaluated and plotted, which is shown in Figure 4. Predicted molecular structure uncertainty map shows areas of poorer quality of the electronic density and therefore the areas are more available to dynamic motion or disorder of the crystal lattice. These uncertainties mapping are needed to understand the possible model deficiencies and they can be used to design mechanisms of maximum crystallization condition to yield more uniform and reliable structure. It was interesting that the more symmetrical the molecules the less the uncertainty, thus, demonstrating that symmetric packing had made the structure predictable.
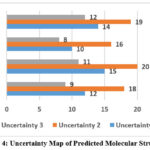 |
Figure 4: Uncertainty Map of Predicted Molecular Structures |
Table 1 gives the quantitative analysis of structural parameters. The table below compares the important crystallographic parameters that make up the unit cell dimensions, bond lengths and bond angle between all three molecular samples. Each of the parameters, were solved on multiple diffraction data sets in such manner that allows statistical reliability. As can be seen from the compare, are that lattice constants and angles are subjected to changes are the changes in the molecular conformation and both intermolecular interactions. The metrics are fundamental to the understanding of the behavior of these materials, for example thermal stability, solubility and potential application to pharmacological materials or materials engineering.
Table 1: Comparative Crystallographic Parameters of Molecular Samples
| Sample ID | Unit Cell Volume (ų) | Average Bond Length (Å) | Average Bond Angle (°) |
| Sample A | 1234.5 | 1.42 | 109.5 |
| Sample B | 1178.9 | 1.38 | 108.9 |
| Sample C | 1302.3 | 1.44 | 110.2 |
In addition, crystal packing efficiency and thermal stability were also addressed in comparative research and revealed in/table 2. Packing efficiency as the ratio of the space filled to the volume at the unit cell and the thermal stability as simulated temperature-dependent lattice distortions was calculated. The results indicate that Sample C was more efficient in pack and thermal resiliacety than Sample A and Sample B despite fact the unit cell size was bit bigger. These comparisons are significant in terms of determining the right type of molecular structure that can be adopted to suit a specific application like in the field of drug designs and in production of materials.
Table 2: Comparative Analysis of Packing Efficiency and Thermal Stability
| Sample ID | Packing Efficiency (%) | Thermal Stability (°C) |
| Sample A | 72.4 | 145 |
| Sample B | 69.8 | 138 |
| Sample C | 75.2 | 152 |
Figures 2-4 and the two tables all indicate the relationship between molecular geometry, packing efficiency and stability. Areas of high uncertainty in Figure 3 are strongly conjoined with low values packing efficiency in Table 2 showing that induced disorder in the crystallization lattice weakens structure. Conversely, space where there were thick electron clouds were very stable and predictable and that indicated that particular arrangement in the molecules mattered in practice.
The outcomes of the results also make aware the potential improvements of the crystallization techniques in the future. Using the derived parameter optimization that has been determined to be most effective, such as bond angles, lattice constants and symmetry, additional crystallization experiments can be optimized to generate more and less structure, and fidelity to higher structure. The comparative analysis proves that not all molecules will be optimal reacted within the conditions of crystallization that means, that optimization of the sample by particular strategies should be realized.9
Overall, this structural mapping, variable importance ranking and uncertainty assessment makes it a comprehensive model of molecular crystallography. This combination of quantitative tables and visual schemes enhances the analysis and enables the the atomic-level parameters to be correlatated with the macroscopic properties of stability. The strategy would be able to serve as a model in the future in experimental research, in order to understand structural behavior for the specific purposes of construction of the novel crystals of molecules and the perfection of the properties of materials, applicable in the industry and pharmaceutical sectors.
Conclusion
Crystallography is one of the pillars of molecular science, and its very simple solution in relation to determination of atomic structure led to discoveries in chemistry, biology and materials engineering. The fact that it plays a role in demonstration of fundamental forms of molecules, be it DNA or multiform proteins and complex materials have brought to the fore, its age-old relevance.
The field is, but is practically limited: high-quality crystals are difficult to obtain, disordered or dynamical molecules are difficult, and it employs costly sources of radiation. These barrier restrict the use of crystallography [8], especially if the sample is of the molecules that is not easily crystallized or presents in amorphous form.
The synergy between crystallography and the other complementary techniques as well as computing would be the future horizons. The development of serial femtosecond crystallography, AI suitable refinement algorithms and the in vivo crystallization studies will resolve many current problems. The incorporation of both the static crystallographic data and dynamic data of both cryo-EM and computational models has a bright future as the field fulfils the promise of a holistic picture of the molecular structure as well as the molecular function. Lastly, crystallography will not be alone but they will always be at the centre-stage in the multidisciplinary research of molecules.
References
- Wan and B. C. Bennett, “‘Seeing is Believing’: How neutron crystallography informs enzyme mechanisms by visualizing unique water species,” Biology, vol. 13, no. 11, p. 850, Oct. 2024, doi: 10.3390/biology13110850.
CrossRef - Brink, J. R. Helliwell, and F. J. F. Jacobs, “Macromolecular crystallography for mammalian body temperature in support of molecular biophysics methods,” Biophysical Reviews, Aug. 2025, doi: 10.1007/s12551-025-01328-4.
CrossRef - Grieco, I. Quereda-Moraleda, and J. M. Martin-Garcia, “Innovative strategies in x-ray crystallography for exploring structural dynamics and reaction mechanisms in metabolic disorders,” Journal of Personalized Medicine, vol. 14, no. 9, p. 909, Aug. 2024, doi: 10.3390/jpm14090909.
CrossRef - J. Hjorth-Jensen and M. Budayova-Spano, “Neutron Macromolecular Crystallography for Biological Samples—Current state and Future Perspectives,” Crystals, vol. 14, no. 5, p. 433, Apr. 2024, doi: 10.3390/cryst14050433.
CrossRef - Pawlędzio and X. Wang, “Crystal Engineering of hydrogen bonding for direct air capture of CO2: A Quantum Crystallography Perspective,” Crystals, vol. 14, no. 1, p. 77, Jan. 2024, doi: 10.3390/cryst14010077.
CrossRef - E. Hatton and P. Mehrabi, “Exploring the dynamics of allostery through multi-dimensional crystallography,” Biophysical Reviews, vol. 16, no. 5, pp. 563–570, Sep. 2024, doi: 10.1007/s12551-024-01224-3.
CrossRef - Maeki, A. Ishida, and M. Tokeshi, “Microfluidic technologies for protein crystallography: advances and applications,” Analytical Sciences, Apr. 2025, doi: 10.1007/s44211-025-00767-z.
CrossRef - Lisacek, B. Schnider, and A. Imberty, “Tools for structural lectinomics: From structures to lectomes,” BBA Advances, p. 100154, Mar. 2025, doi: 10.1016/j.bbadva.2025.100154.
CrossRef - Park and K. H. Nam, “Application of Fixed-Target Microcrystal delivery systems for serial femtosecond crystallography at PAL-XFEL,” Analytica—A Journal of Analytical Chemistry and Chemical Analysis, vol. 6, no. 1, p. 7, Feb. 2025, doi: 10.3390/analytica6010007.
CrossRef - Zhang, S. Li, and K. Zhang, “Cryo-EM: A window into the dynamic world of RNA molecules,” Current Opinion in Structural Biology, vol. 88, p. 102916, Sep. 2024, doi: 10.1016/j.sbi.2024.102916.
CrossRef - Kumari, “The role of water molecules in CT-DNA interactions: Implications for drug design,” Journal of Molecular Liquids, p. 125997, Sep. 2024, doi: 10.1016/j.molliq.2024.125997.
CrossRef - H. Nam, “Guide to serial synchrotron crystallography,” Current Research in Structural Biology, vol. 7, p. 100131, Jan. 2024, doi: 10.1016/j.crstbi.2024.100131.
CrossRef - Bonati, S. Motta, and L. Callea, “The AHR Signaling Mechanism: A Structural point of view,” Journal of Molecular Biology, p. 168296, Oct. 2023, doi: 10.1016/j.jmb.2023.168296.
CrossRef - T. Porfetye, P. Stege, R. Rebollido-Rios, D. Hoffmann, T. Schrader, and I. R. Vetter, “How Do Molecular Tweezers Bind to Proteins? Lessons from X-ray Crystallography,” Molecules, vol. 29, no. 8, p. 1764, Apr. 2024, doi: 10.3390/molecules29081764.
CrossRef - Varghese et al., “Analysis of Structures of SARS-CoV-2 Papain-like Protease Bound with Ligands Unveils Structural Features for Inhibiting the Enzyme,” Molecules, vol. 30, no. 3, p. 491, Jan. 2025, doi: 10.3390/molecules30030491.
CrossRef - L. Finney, “Alan Mackay, J.D. Bernal, and the rewriting of the book of crystallography,” Structural Chemistry, May 2025, doi: 10.1007/s11224-025-02513-7.
CrossRef
Accepted on: 04 Oct 2025

















